Research Article
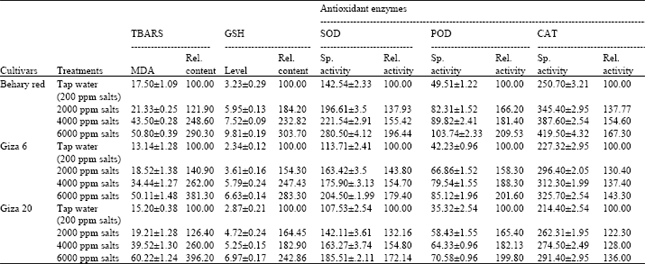
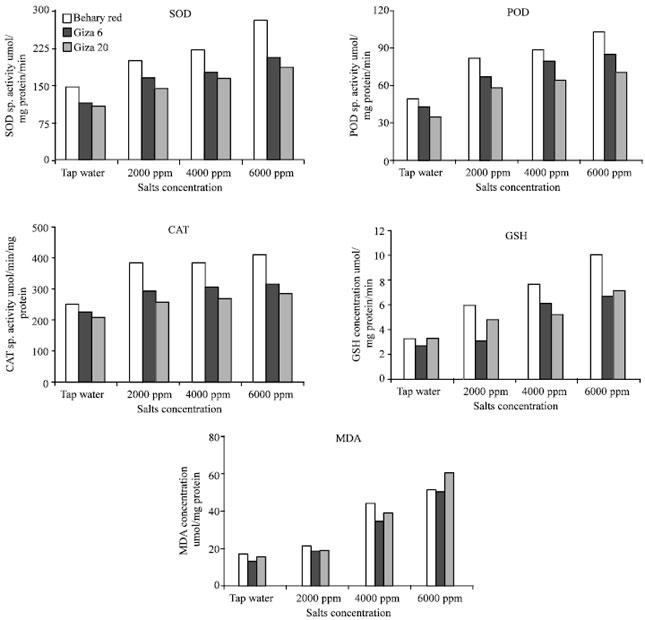
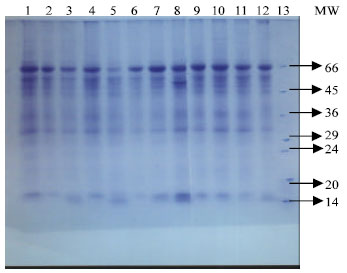
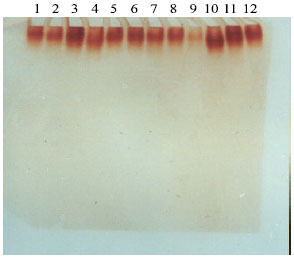
Influence of Salinity on Lipid Peroxidation, Antioxidant Enzymes and Electrophoretic Patterns of Protein and Isoenzymes in Leaves of Some Onion Cultivars
Not Available
H. Hanaa
Not Available
Mohamed, A. Amal
Not Available
M.M. Hussein
Not Available