Research Article
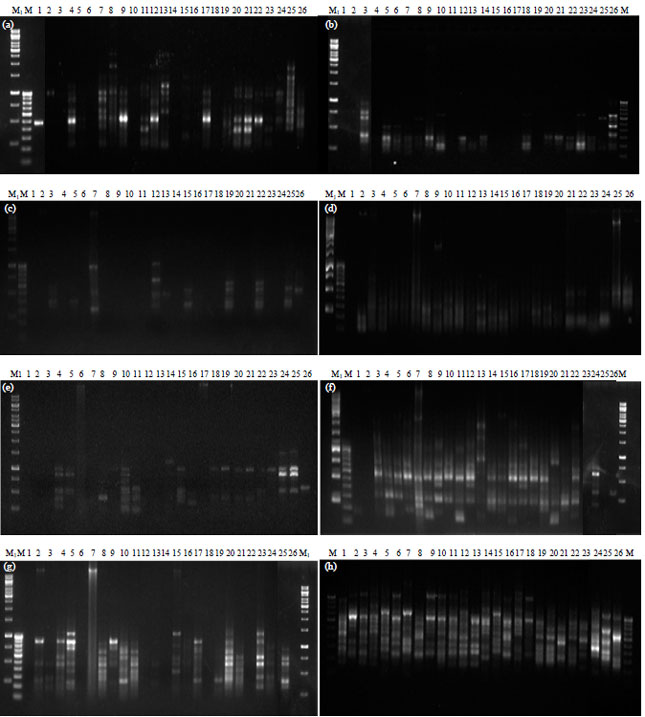
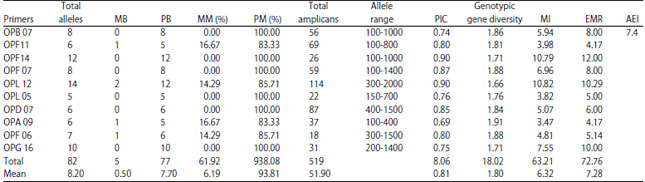
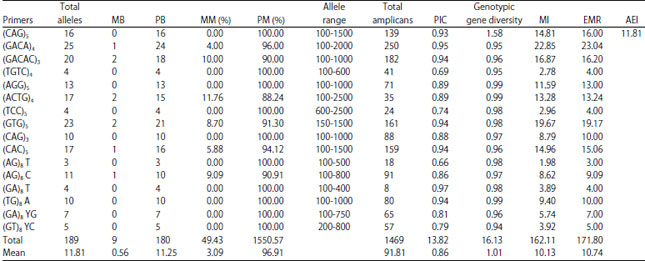
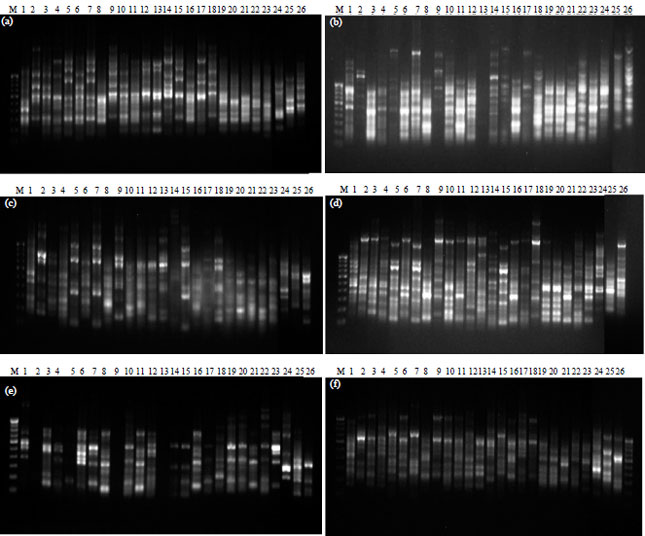
Comparative Assessment of RAPD and ISSR Markers to Study Genetic Polymorphism in Colletotrichum gloeosorioides Isolates of Mango
Department of Plant Pathology, Centre for Plant Protection Studies, Tamil Nadu Agricultural University, 641003 Coimbatore, Tamil Nadu, India
LiveDNA: 91.16234
Prabukarthikeyan Karthikeyan Rathinam
Division of Crop Protection, National Rice Research Institute (NRRI), Cuttack, Odisha, India
Raguchander Thiruvengadum
Department of Plant Pathology, Centre for Plant Protection Studies, Tamil Nadu Agricultural University, 641003 Coimbatore, Tamil Nadu, India
Prabakar Kuppusamy
Department of Plant Pathology, Centre for Plant Protection Studies, Tamil Nadu Agricultural University, 641003 Coimbatore, Tamil Nadu, India