ABSTRACT
The Q gene of wheat is a member of the AP2 class of transcription factors and has been shown to influence numerous morphological and domestication-related characters including plant height, flowering time, leaf morphology and spike architecture. To identify candidate genes involved with the transcriptional regulation networks of Q gene signaling pathways, I used the yeast two-hybrid system to identify proteins that directly interact with the Q protein. Five potential Q interactors were identified and they include protein kinases, transcription factors and a stress responsive protein. This research provides the basis for unraveling the signaling pathways and genetic networks governed by the Q gene.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajpp.2007.295.302
URL: https://scialert.net/abstract/?doi=ajpp.2007.295.302
INTRODUCTION
Wheat, including bread wheat (Triticum aestivum L., 2n = 6x = 42, AABBDD genomes) and durum wheat (T. turgidum L., 2n = 4x = 28, AABB genomes), is one of three major cereal crops consumed throughout the world. The wheat Q gene pleiotropically affects characteristics, such as free threshing, heading time, plant height, spike length, spikelet size, seed fertility, rachis fragility and glume shape and tenacity (Muramatsu, 1986; Kato et al., 2003; Jantasuriyarat et al., 2004; Simons et al., 2006). Thus, Q is a major regulatory gene involved in both vegetative and floral development. The Q gene was recently cloned and found to belong to the AP2/ERF family of transcription factors (Simons et al., 2006).
The AP2/ERF family of transcription factors consists of three subfamilies (Riechmann et al., 2000). The RAV subfamily is characterized by one AP2 DNA-binding domain and a B3 DNA-binding domain. They are involved in ethylene response (Alonso et al., 2003) and brassinosteroid response (Hu et al., 2004). The ERF subfamily has one AP2 binding domain and its members have very diverse functions including hormonal signal transduction (Ohme-Takagi and Shinshi, 1995), biotic and abiotic stress response (Stockinger et al., 1997; Liu et al., 1998; Dubouzet et al., 2003), metabolism regulation (van der Fits and Memelink, 2000; Aharoni et al., 2004; Broun et al., 2004; Zhang et al., 2005) and developmental functions (van der Graaff et al., 2000; Banno et al., 2001; Chuck et al., 2002). The third subfamily is the AP2 subfamily having two AP2 binding domains and regulates various developmental processes. The Q gene is classified into this subfamily due to the presence of two AP2 DNA-binding domains (Simons et al., 2006).
Fourteen genes have been classified as belonging to the AP2 subfamily in Arabidopsis (Kim et al., 2006). Of these, APETALA2 (AP2) has been the most extensively characterized. AP2 is involved in establishing floral meristem identity and regulating floral homeotic gene expression (Komaki et al., 1988; Bowman et al., 1991; Jofuku et al., 1994). AP2 also regulates the stem cell niche in shoot meristems in a dosage dependent manner (Wurschum et al., 2006) and it influences gibberellic regulation of metabolism in seed development, sucrose sensing, flowering time, leaf number, soluble sucrose metabolism and yield (Jofuku et al., 2005; Ohto et al., 2005). Analysis of mutants and/or over expression studies of other Arabidopsis genes within the AP2 subfamily, such as TOE1, TOE2, WRINKLED1, SCHLAFUMTZE, SCHNARCHZAPFEN, ANTEGUMENTA, PLETHORA1, PLETHORA2 and BABYBOOM, have implicated them in developmental roles (Boutilier et al., 2002; Aukerman and Sakai, 2003; Schmid et al., 2003; Aida et al., 2004; Cernac and Benning, 2004; Nole-Wilson et al., 2005).
Outside of Arabidopsis, the study of AP2-like genes has been very limited. In maize INDETERMINATE SPIKELET1 suppresses indeterminate growth of the spikelet (Chuck et al., 1998) and GLOSSY15 regulates the identity of leaf epidermal cells during the vegetative phase transition from juvenile to adult plants (Moose and Sisco, 1996). Other AP2-like genes have been identified in other flowering plants (Maes et al., 2001; Boutilier et al., 2002) and gymnosperms (Vahala et al., 2001), but thus far no genes within the AP2 subfamily have been reported to have functions other than developmental regulation. Yeast two-hybrid (Y2H) experiments were used to begin unraveling the signaling network associated with Q.
MATERIALS AND METHODS
Plant Materials
The Q gene cDNA was cloned from leaves of T. aestivum cultivar Chinese Spring (CS). Leaf tissue of the T. durum cultivar Langdon (LDN) was used to construct a cDNA library for yeast two-hybrid screening. Spike tissue of T. aestivum cultivar Bobwhite (QQ genotype) was used for the cloning of genes in the targeted Y2H approach.
Plasmid Vectors, Bacterial and Yeast Strains and Culture Conditions
E. coli strains DH5α and TOP10 (Invitrogen, Carlsbad, CA, USA) were used for transformation after DNA ligation and TOPO cloning, respectively. Constructs in the bait vector pGBKT7 were selected by 50 μg mL-1 kanamycin and constructs in the prey vector pGADT7 were under 200 μg mL-1 ampicillin selection. All E. coli cells were grown in standard Luria-Bertrani medium at 37°C.
Cloning of Full-length cDNAs of Q and Putative Q Interactors
Wheat total RNA was isolated using TRIzol® Reagent (Invitrogen). To clone the full-length coding region of the Q gene, reverse transcription polymerase chain reaction (RT-PCR) was performed according to the protocol of the SMARTTM RACE cDNA Amplification kit (Clontech, Palo Alto, CA, USA) followed by PCR amplification using primers 5’-CATATG GTG CTG GAT CTC AAT GTG GAG TC-3’ (created NdeI restriction enzyme site underlined) and 5’-GGATCC TCA GTT GTC CGG CGG GCG GGG GAA-3’ (created BamHI site underlined) based on GenBank sequence AY702956. The cloned Q gene coding region was excised from the vector by NdeI and BamHI restriction enzymes and ligated into pGBKT7 as the bait for yeast two-hybrid analysis. For the Q interactors, first-strand cDNA was synthesized from total RNA using a random hexamer primer followed by PCR amplification using the primers listed in Table 1. PCR amplifications were performed using a combination of Taq DNA polymerase (New England Biolabs, Beverly, MA, USA) and Pfu Turbo (Stratagene, La Jolla, CA, USA) with the GeneAmp PCR System 9700 (ABI, Applied Biosystems, Foster City, CA, USA). PCR products were cloned into pCR® 2.1-TOPO® vector (Invitrogen) and sequenced. Recombinant plasmid DNA was isolated using the QIAGEN (Valencia, CA, USA) mini-prep kit.
Yeast Two-hybrid Analyses for Identification of Q Interactors
The yeast two-hybrid MatchmakerTM Library Construction and Screening kit was used for analysis of protein-protein interactions according to the manufacturer’s protocols (Clontech). The Q gene was ligated into pGBKT7 as bait and pGADT7 was used to construct prey of a cDNA library or was used in the targeted Y2H approach. Cells of yeast strain Y187 were transformed with the bait plasmids and AH109 cells were used for the prey plasmids. The Fast Yeast Transformation KitTM (G-Biosciences/Genotech, St. Louis, MO, USA) was used, according to the manufacturer’s protocols.
| Table 1: | Primers used to clone putative wheat Q interactors. Restriction sites are underlined |
 | |
For Y2H library screening, a cDNA library was constructed in pGADT7, according to the instructions provided by the manufacturer (Clontech). Total RNA from leaves of wheat seedlings at the 2-leaf-stage was used as template for the cDNA library construction. For targeted Y2H analysis, cDNAs were generated from total RNA isolated from immature spikes of wheat. Yeast strains Y187 and AH109 were mated on YPD plates overnight. Then the yeast cells were cultured in SD medium without leucine (L) and tryptophan (W) for two days. To test the protein interactions, the SD/-L-W cultured yeast cells were inoculated on SD plates containing X-Gal but lacking leucine, tryptophan, histidine (H) and adenine (A). The empty vectors pGBKT7 and pGADT7 were used as negative controls and the interaction between p53 and LTA (large T-antigen) served as a positive control. Prey plasmids were isolated from yeast and transformed into E. coli. The isolated plasmid from E. coli was then retransformed back to yeast AH109 and the mating assays were repeated using SD medium with X-Gal and under -L-W-H-A stringent selection. The Q interactors of positive colonies from the cDNA library screening were sequenced. Sequence data were queried onto the NCBI database (http://www.ncbi.nlm.nih.gov/) and the DFCI (http://compbio.dfci.harvard.edu/tgi/) Triticum aestivum Gene Index (TaGI) using BlastN.
RESULTS AND DISCUSSION
The Q gene is present in all domesticated wheat and it most profoundly affects spike architecture and grain threshability (Muramatsu, 1986). However, the Q gene is also expressed in other tissues, such as roots and leaves and it is known to influence morphology of the vegetative tissues (Simons et al., 2006). Whereas members of the AP2 subfamily of AP2 genes have not been associated with stress, many members of the other AP2 subfamilies are stress-related genes. Therefore, I investigated the role of the Q gene in wheat in response to abiotic stresses.
Because the Q gene product is a transcription factor, I hypothesized that protein-protein interactions play a critical role for the regulation of transcriptional activity. Q is not auto-activated in the Y2H system, as confirmed by the absence of interaction with AD empty vector or LTA (Fig. 1). Similar to most transcription factors, Q has the ability to form a homodimer (Fig. 1). To further understand how the Q gene is involved in regulatory networks, I screened a cDNA library and also applied a targeted Y2H approach to identify gene products that directly interact with the Q protein.
After screening approximately 10 million yeast colonies from a cDNA library made from leaves, 52 initial putative positive clones were further validated by a second round of selection. Three independent clones of the same gene were confirmed to be positive for encoding a Stress Responsive Protein (SRP) (Table 2, Fig. 2). Interestingly, a rice homolog of the stress responsive protein was characterized to be induced by ABA and salt (Moons et al., 1995). The exact biochemical function of this stress responsive protein remains unknown.
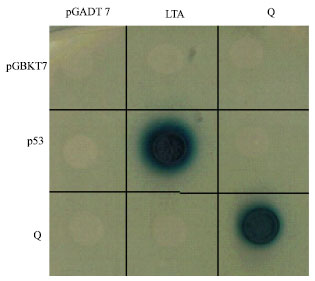 | |
| Fig. 1: | Yeast two-hybrid analysis. Bait: pGBKT7 (empty vector), p53 and Q. Prey: pGADT7 (empty vector), LTA (large T-antigen) and Q. The interaction between p53 and LTA serves as a positive control. Yeast cells were incubated under highly stringent selection (the absence of leucine, tryptophan, histidine and adenine) and the medium contained X-Gal. The yeast cells can grow and show blue on the plate only when the bait and prey proteins interact with each other |
Furthermore, I followed the targeted Y2H approach published by Menke et al. (2005), especially for the low expression levels of transcription factors. Because the Q protein is a member of the AP2 transcription factor family and is physiologically responsive to stresses, I hypothesize that the Q protein may interact with other proteins that are involved in stress stimuli. Therefore, I chose well-characterized transcription factors (ERF, TGA and WRKY) and protein kinases (SOS2 and PKS3) as candidates.
I did identify an ERF in wheat that interacted with the Q protein, while there were no positive interactions between any WRKY protein and the Q protein. The AP2/ERF family of transcription factors includes the ERF and AP2 subfamilies (Riechmann et al., 2000; McGrath et al., 2005; Nakano et al., 2006). AP2/ERF proteins are important for the transcriptional regulation of development and various responses to environmental stimuli (Riechmann and Meyerowitz, 1998). The ERF family proteins usually participate in the crosstalk of regulatory networks in response to different stimuli, such as salt, drought, cold and ABA (Shinozaki et al., 2003; Yamaguchi-Shinozaki and Shinozaki, 2005; Fujita et al., 2006). I designated the wheat ERF-like protein that interacted with the Q protein as TaERF7 because of its similarity to AtERF7 (GenBank accession number NP-188666), which was found to have 38% identity at the amino acid level. In Arabidopsis, AtERF7 plays a critical role in drought and ABA responses and the transcriptional complex may be regulated by PKS3 (Song et al., 2005), which also interacts with the Q protein. AtERF7 interacts with and is thereby phosphorylated by PKS3. The regulation of the AtERF7 network may involve chromatin remodeling, because AtERF7 also interacts with AtSin3 that in turn interacts with the histone deacetylase HDA19 (Song et al., 2005).
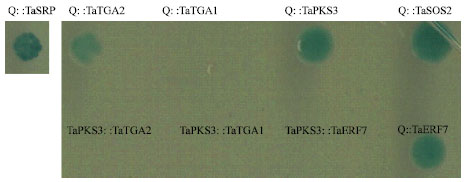 | |
| Fig. 2: | Putative wheat Q interactors identified by library screening and the direct targeted approach. Each pair is shown as bait::prey combination. The selection condition was the same as in Fig. 1 and also described in the Materials and Methods |
| Table 2: | Identified wheat Q interactors by yeast two-hybrid analysis |
 | |
Intriguingly, I also identified PKS3 and SOS2 homologs in wheat, designated TaPKS3 and TaSOS2, respectively, which interacted with the Q protein. Both SOS2 and PKS3 are Ser/Thr protein kinases. Arabidopsis SOS2 interacts with another protein kinase known as SOS3 and plays a role in salt tolerance (Guo et al., 2004). PKS3 physically interacts with the protein phosphatase ABI2 and is a negative regulator of ABA responses (Guo et al., 2002). SOS2 only weakly interacts with AtERF7, but PKS3 strongly interacts with AtERF7 (Song et al., 2005). Whereas both TaSOS2 and TaPKS3 can interact with Q protein, no detectable interaction was identified between TaPKS3 and TaERF7 (Fig. 2).
The TGA family of transcription factors belongs to a class of basic leucine zipper (bZIP) proteins. NPR1 is a critical regulator of the SA-mediated systemic acquired resistance (Cao et al., 1994; Delaney et al., 1995). It has been shown in Arabidoposis that members of the TGA family proteins differentially interact with NPR1 in the Y2H system, with TGA2, TGA3 and TGA6 showing strong binding (Zhang et al., 1999; Zhou et al., 2000), whereas TGA1 and TGA4 do not bind to NPR1 in Y2H assays. These results show that the Q protein also interacts differentially with TGA transcription factors because Q interacted with TaTGA2 but not with TaTGA1 (Fig. 2). In addition to the results of Arabidopsis, accumulating evidence suggests that NPR1 is pivotal for the regulation network in response to pathogens in rice (Chern et al., 2005) and wheat (Makandar et al., 2006). Usually there are cross-talking networks among responses to ABA, JA, SA and ethylene (Pieterse and Van Loon, 2004; Fujita et al., 2006). How the Q protein integrates the signaling pathways is largely unknown so far. However, the identification of Q interactors, including protein kinases and transcription factors, provides the basis for unraveling the pathways and mechanisms of Q-mediated regulation.
ACKNOWLEDGMENTS
The author thanks T.L. Tranby, D. Holmes, S. Xu, K. Simons and J. Faris. This research was funded by USDA-ARS CRIS project No. 5442-21000-030-00D. I feel deeply grateful to Dr. Brady Vick and Dr. Lynn Dahleen for pivotal comments on this manuscript.
REFERENCES
- Aharoni, A., S. Dixit, R. Jetter, E. Thoenes, G. van Arket and A. Pereira, 2004. The SHINE clade of AP2 domain transcription factors activates wax biosynthesis, alters cuticle properties and confers drought tolerance when overexpressed in Arabidopsis. Plant Cell, 16: 2463-2480.
CrossRefDirect Link - Aida, M., D. Beis, R. Heidstra, V. Willemsen and I. Blilou et al., 2004. The PLETHORA genes mediate patterning of the Arabidopsis root stem cell niche. Cell, 119: 109-120.
Direct Link - Alonso, J.M., A.N. Stepanova, T.J. Leisse, C.J. Kim and H. Chen et al., 2003. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science, 301: 653-657.
CrossRefPubMedDirect Link - Aukerman, M. and H. Sakai, 2003. Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell, 15: 2730-2741.
Direct Link - Banno, H., Y. Ikeda, Q.W. Niu and N.H. Chua, 2001. Overexpression of Arabidopsis ESR1 induces initiation of shoot regeneration. Plant Cell, 13: 2609-2618.
Direct Link - Boutilier, K., R. Offringa, V.K. Sharma, H. Kieft and T. Ouellet et al., 2002. Ectopic expression of BABY BOOM triggers a conversion from vegetative to embryonic growth. Plant Cell, 14: 1737-1749.
CrossRefDirect Link - Bowman, J.L., D.R. Smyth and E.M. Meyerowitz, 1991. Genetic interactions among floral homeotic genes of Arabidopsis. Development, 112: 1-20.
PubMedDirect Link - Broun, P., P. Poindexter, E. Osborne, C.Z. Jiang and J.L. Riechmann, 2004. WIN1, a transcriptional activator of epidermal wax accumulation in Arabidopsis. Proc. Natl. Acad. Aci. USA., 101: 4706-4711.
CrossRefPubMedDirect Link - Cao, H., S.A. Bowling, A.S. Gordon and X. Dong, 1994. Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell, 6: 1583-1592.
Direct Link - Cernac, A. and C. Benning, 2004. WRINKLED encodes an AP2/EREB domain protein involved in the control of storage compound biosynthesis in Arabidopsis. Plant J., 40: 575-585.
Direct Link - Chern, M., P.E. Canlas, H.A. Fitzgerald and P.C. Ronald, 2005. Rice NRR, a negative regulator of disease resistance, interacts with Arabidopsis NPR1 and rice NH1. Plant J., 43: 623-635.
Direct Link - Chuck, G., R.B. Meeley and S. Hake, 1998. The control of maize spikelet meristem fate by the APETALA2-like gene Indeterminate spikelet1. Genes. Dev., 12: 1145-1154.
Direct Link - Chuck, G., M. Muszynski, E. Kellogg, S. Hake and R.J. Schmidt, 2002. The control of spikelet meristem identity by the Branched silkless1 gene in maize. Science, 298: 1238-1241.
Direct Link - Delaney, T.P., L. Friedrich and J.A. Ryals, 1995. Arabidopsis signal transduction mutant defective in chemically and biologically induced disease resistance. Proc. Natl. Acad. Sci. USA., 92: 6602-6606.
Direct Link - Dubouzet, J.G., Y. Sakuma, Y. Ito, M. Kasuga and E.G. Dubouzet et al., 2003. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought, high-salt and cold-responsive gene expression. Plant J., 33: 751-763.
CrossRefDirect Link - Fujita, M., Y. Fujita, Y. Noutoshi, F. Takahashi, Y. Narusaka, K. Yamaguchi-Shinozaki and K. Shinozaki, 2006. Crosstalk between abiotic and biotic stress responses: A current view from the points of convergence in the stress signaling networks. Curr. Opin. Plant Biol., 9: 436-442.
Direct Link - Guo, Y., L. Xiong, C.P. Song, D. Gong, U. Halfter and J.K. Zhu, 2002. A calcium sensor and its interacting protein kinase are global regulators of abscisic acid signaling in Arabidopsis. Dev. Cell, 3: 233-244.
Direct Link - Guo, Y., Q.S. Qiu, F.J. Quintero, J.M. Pardo and M. Ohta et al., 2004. Transgenic evaluation of activated mutant alleles of SOS2 reveals a critical requirement for its kinase activity and C-terminal regulatory domain for salt tolerance in Arabidopsis thaliana. Plant Cell, 16: 435-449.
Direct Link - Hu, Y.X., Y.X. Wang, X.F. Liu and J.Y. Li, 2004. Arabidopsis RAV1 is down-regulated by brassinosteroid and may act as a negative regulator during plant development. Cell Res., 14: 8-15.
PubMedDirect Link - Jantasuriyarat, C., M.I. Vales, C.J.W. Watson and O. Riera-Lizarazu, 2004. Identification and mapping of genetic loci affecting the free-threshing habit and spike compactness in wheat (Triticum aestivum L.). Theor. Applied Genet., 108: 261-273.
Direct Link - Jofuku, K.D., B.G.W. den Boer, M. van Montagu and J.K. Okamuro, 1994. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell, 6: 1211-1225.
Direct Link - Jofuku, K.D., P.K. Omidyar, Z. Gee and J.K. Okamuro, 2005. Control of seed mass and seed yield by the floral homeotic gene APETALA2. Proc. Nat. Acad. Sci. USA., 102: 3117-3122.
Direct Link - Kato, K., R. Sonokawa, H. Miura and S. Sawada, 2003. Dwarfing effect associated with the threshability gene Q on wheat chromosome 5A. Plant Breed., 122: 489-492.
Direct Link - Kim, S., P.S. Soltis, K. Wall and D.E. Soltis, 2006. Phylogeny and domain evolution in the APETALA2-like gene family. Mol. Biol. Evol., 23: 107-120.
CrossRefDirect Link - Liu, Q., M. Kasuga, Y. Sakuma, H. Abe, S. Miura, K. Yamaguchi-Shinozaki and K. Shinozaki, 1998. Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell, 10: 1391-1406.
Direct Link - Maes, T., N. Van de Steene, J. Zethof, M. Karimi and M. D-Hauq et al., 2001. Petunia Ap2-like genes and their role in flower and seed development. Plant Cell, 13: 229-244.
Direct Link - Makandar, R., J.S. Essig, M.A. Schapaugh, H.N. Trick and J. Shah, 2006. Genetically engineered resistance to Fusarium head blight in wheat by expression of Arabidopsis NPR1. Mol. Plant Microbe Interact., 19: 123-129.
Direct Link - McGrath, K.C., B. Dombrecht, J.M. Manners, P.M. Schenk and C.I. Edgar et al., 2005. Repressor and activator-type ethylene response factors functioning in jasmonate signaling and disease resistance identified via a genome-wide screen of Arabidopsis transcription factor gene expression. Plant Physiol., 139: 949-959.
Direct Link - Menke, F.L.H., H.G. Kang, Z. Chen, J.M. Park, D. Kumar and D.F. Klessig, 2005. Tobacco transcription factor WRKY1 is phosphorylated by the MAP kinase SIPK and mediates HR-like cell death in tobacco. Mol. Plant Microbe Interact., 18: 1027-1034.
Direct Link - Moons, A., G. Bauw, E. Prinsen, M. van Montagu and D. van der Straeten, 1995. Molecular and physiological responses to abscisic acid and salts in roots of salt-sensitive and salt-tolerant Indica rice varieties. Plant Physiol., 107: 177-186.
CrossRef - Moose, S.P. and P.H. Sisco, 1996. Glossy15, an APETALA2-like gene from maize that regulates leaf epidermal cell identity. Genes Dev., 10: 3018-3027.
Direct Link - Nakano, T., K. Suzuki, T. Fujimura and H. Shinshi, 2006. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol., 140: 411-432.
Direct Link - Nole-Wilson, S., T.L. Tranby and B.A. Krizek, 2005. AINTEGUMENTA-like (AIL) genes are expressed in young tissues and may specify meristematic or division-competent states. Plant Mol. Biol., 57: 613-628.
Direct Link - Ohme-Takagi, M. and H. Shinshi, 1995. Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell, 7: 173-182.
Direct Link - Ohto, M., R.L. Fischer, R.B. Goldberg, K. Nakamura and J.J. Harada, 2005. Control of seed mass by APETALA2. Proc. Natl. Acad. Sci. USA., 102: 3123-3128.
Direct Link - Pieterse, C.M. and L.C. Van Loon, 2004. NPR1: The spider in the web of induced resistance signaling pathways. Curr. Opin. Plant Biol., 7: 456-464.
Direct Link - Riechmann, J.L. and E.M. Meyerowitz, 1998. The AP2/EREBP family of plant transcription factors. Biol. Chem., 379: 633-646.
Direct Link - Riechmann, J.L., J. Heard, G. Martin, L. Reuber and C.Z. Jiang et al., 2000. Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science, 290: 2105-2110.
CrossRefPubMedDirect Link - Schmid, M., N.H. Uhlenhaut, F. Godard, M. Demar, R. Bressan, D. Weigel and J.U. Lohmann, 2003. Dissection of floral induction pathways using global expression analysis. Development, 130: 6001-6012.
Direct Link - Shinozaki, K., K. Yamaguchi-Shinozaki and M. Seki, 2003. Regulatory network of gene expression in the drought and cold stress responses. Curr. Opin. Plant Biol., 6: 410-417.
PubMedDirect Link - Simons, K.J., J.P. Fellers, H.N. Trick, Z. Zhang, Y.S. Tai, B.S. Gill and J.D. Faris, 2006. Molecular characterization of the major wheat domestication gene Q. Genetics, 172: 547-555.
Direct Link - Song, C.P., M. Agarwal, M. Ohta, Y. Guo, U. Halfter, P. Wang and J.K. Zhu, 2005. Role of an Arabidopsis AP2/EREBP-type transcriptional repressor in abscisic acid and drought stress responses. Plant Cell, 17: 2384-2396.
Direct Link - Stockinger, E.J., S.J. Gilmour and M.F. Thomashow, 1997. Arabidopsis thaliana CBF1 encodes an AP2 domain-containing transcriptional activator that binds to the C-repeat/DRE, a cis-acting DNA regulatory element that stimulates transcription in response to low temperature and water deficit. Proc Natl. Acad. Sci. USA., 94: 1035-1040.
Direct Link - Vahala, T., B. Oxelman and S. von Arnold, 2001. Two APETALA2-like genes of Picea abies are differentially expressed during development. J. Exp. Bot., 52: 1111-1115.
Direct Link - Van der Fits, L. and J. Memelink, 2000. ORCA3, a jasmonate-responsive transcriptional regulator of plant primary and secondary metabolism. Science, 289: 295-297.
CrossRefDirect Link - van der Graaff, E., A.D. Dulk-Ras, P.J. Hooykaas and B. Keller, 2000. Activation tagging of the LEAFY PETIOLE gene affects leaf petiole development in Arabidopsis thaliana. Development, 127: 4971-4980.
Direct Link - Wurschum, T., R. Gross-Hardt and T. Laux, 2006. APETALA2 regulates the stem cell niche in the Arabidopsis shoot meristem. Plant Cell, 18: 295-307.
Direct Link - Yamaguchi-Shinozaki, K. and K. Shinozaki, 2005. Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci., 10: 88-94.
CrossRefDirect Link - Zhang, Y., W. Fan, M. Kinkema, X. Li and X. Dong, 1999. Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene. Proc. Natl. Acad. Sci. USA., 96: 6523-6528.
Direct Link - Zhang, J.Y., C.D. Broeckling, E.B. Blancaflor, M.K. Sledge, L.W. Sumner and Z.Y. Wang, 2005. Overexpression of WXP1, a putative Medicago truncatula AP2 domain-containing transcription factor gene, increases cuticular wax accumulation and enhances drought tolerance in transgenic alfalfa (Medicago sativa). Plant J., 42: 689-707.
Direct Link - Zhou, J.M., Y. Trifa, H. Silva, D. Pontier, E. Lam, J. Shah and D.F. Klessig, 2000. NPR1 differentially interacts with members of the TGA/OBF family of transcription factors that bind an element of the PR-1 gene required for induction by salicylic acid. Mol. Plant Microbe Interact., 13: 191-202.
Direct Link








