ABSTRACT
Drought is a pervasive problem throughout most of the world's cultivated regions and it has been recognized as the single most important limitation to productivity. The present study was carried out to evaluate two maize inbred lines for their drought stress tolerance in the field. These two inbred lines were chosen as drought tolerant (Sd-63) and drought sensitive (Gm-18) for hybridization to obtain the F1 generation and then selfed to obtain the F2 generation. These two maize inbred lines and their F1 and bulks of the two extreme F2 plant groups (the most tolerant F2 group and the most sensitive F2 group) were tested against six RAPD (Random Amplified Polymorphic DNA) primers to determine some molecular markers for drought tolerance in maize. This analysis revealed that, four RAPD primers (A01, A05, A06 and B08) out of the six developed molecular markers for drought tolerance in maize. These RAPD markers could be considered as reliable molecular markers associated with drought tolerance in maize that can be utilized during breeding programs via marker-assisted selection.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajcs.2013.312.318
URL: https://scialert.net/abstract/?doi=ajcs.2013.312.318
INTRODUCTION
Abiotic stress is defined as any change in environmental conditions that might reduce or adversely affect plant growth or development (Bruce et al., 2002). Drought is a pervasive problem throughout most of the world's cultivated regions and it has been recognized as the single most important limitation to productivity. Previous studies indicated that prolonging irrigation intervals led to decreased growth, yield and yield components of maize (De Souza et al., 2009; Shiri et al., 2010; Abdel-Latif et al., 2011). The utilization of molecular markers, developed by analysis of Randomly Amplified Polymorphic DNA (RAPD), for the improvement of drought tolerance is an important part of the solution to improve global maize production. Bulked segregant analysis of F2 plants was developed by Michelmore et al. (1991) as a simpler alternative to isogenic line analysis where the highest and lowest extremes of the F2 population are bulked for the development of RAPD molecular markers needed for QTLs-assisted selection. Several investigators (Abdel-Tawab et al., 2002; Younis et al., 2007) tried to identify molecular markers associated with drought and salt tolerance in maize and sorghum. Therefore, this study aims to identify molecular markers associated with maize drought tolerance via RAPD technique.
MATERIALS AND METHODS
Field experiments: The present study was conducted at the Agricultural Research Station of Ain Shams University, Shalakan, Kalubia Governorate, Egypt, during the three successive growing seasons of 2009, 2010 and 2011.
Two inbred lines of maize (Zea mays, L.), i.e., Sd-63 (drought tolerant) and Gm-18 (drought sensitive), are chosen based on previous screening experiment (Abdel-Latif et al., 2011), Seeds of the two pure lines were obtained from Maize Res. Section, Field Crop Res. Institute, ARC, Giza, Egypt. In 2009 growing season, The grains of the two inbred lines were grown in the field and crossed to obtain the F1 hybrid grains. In 2010 growing season, some of the F1 grains were sown in the field and plants were selfed to obtain the F2 grains. In 2011 growing season, the two maize parental lines and their respective F1 hybrids were grown and evaluated in the field under normal and drought stress conditions. Two separate field trails were carried out; one trail under normal irrigation (irrigation every 13 days) and the other under drought stress (skipping the 4th and 5th irrigations through growing season). The drought stress trail involved the two parental lines, F1 and F2 plants. Each experiment was conducted in a randomized complete block design with three replicates to study the effect of water stress on yield of different maize genotypes. Each replicate consisted of 25 ridges for drought condition and 9 ridges for normal condition. Three ridges were planted for each of P1, P2 and F1 and 16 ridges for F2 under drought condition. Three ridges were planted from each of P1, P2 and F1 under normal irrigation. Each ridge was five meter long and 70 cm width. Planting was done in hills spaced at 25 cm apart and hills were thinned at one plant per hill after about 21 days from sowing. Plant height (cm), 100 kernel weight (g) and grain yield per plant (g) were recorded on all guarded plants in the middle row for each of the P1, P2 and F1 and all guarded plants in all rows of F2 for each replicate. The F2 generation was represented by 700 individual plants. The common agricultural practices of growing maize were applied properly as recommended in the district.
Statistical analysis: The analysis of variance was performed according to the method described by Snedecor and Cochran (1981).
Molecular genetic studies
Genomic DNA extraction: Genomic DNAs were isolated on a small scale from 200 mg of one week old etiolated seedlings of both inbred lines along with their F1 and the two F2 extreme groups (the most drought tolerance and the most drought sensitive group). Leaves were ground to a powder using liquid nitrogen in Eppendorf tubes and DNA were isolated using plant genomic DNA Mini Prep Kit (V-gene Biotechnology, china, Cat. No. 69104) according to the manufacturer manual.
PCR conditions and electrophoresis: Six out of 20 primers for RAPD were used in this study. Names and sequence of the selected primers are illustrated in Table 1.
The amplification conditions and PCR mixture were set according to Williams et al. (1990) for RAPD analysis.
To visualize the PCR products, a 15 μL of each reaction was loaded on 1.2% agarose gels. These gels were run at 90 v for 1 h, visualized with UV Transilluminator and photographed using UVP gel documentation system (Gel Works 1 D advanced software, UVP (Ultraviolet Products).
| Table 1: | Names and sequence of the selected primers for PCR-RAPD |
 | |
Data analysis: Data for each analysis was scored using the UVP gel documentation system. Fragment sizes were estimated using 100 and 1 kb DNA standards (Bioron, Germany).
RESULTS AND DISCUSSION
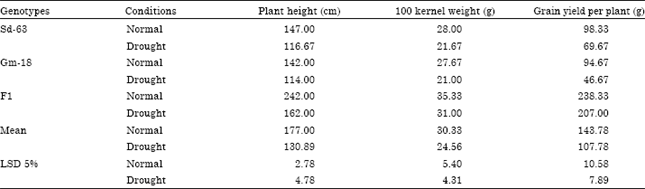
Data for plant height, 100 kernel weight and grain yield per plant for the two maize inbred lines (Sd-63 and Gm-18) with their F1 hybrid under normal and drought condition are presented in Table 2. Significant differences were detected between parental lines for all agronomic traits under normal and water stress conditions indicating the variability existed between the two maize parental lines. Abdel-Sattar and Ahmed (2004), De Souza et al. (2009), Shiri et al. (2010) and Abdel-Latif et al. (2011), in their drought stress experiments on maize, indicated the high genetic potentiality of some maize genotypes based on some agronomic traits.
The results revealed that the parental line Sd-63 was taller (with the means of 147 and 116.67 cm under normal and drought stress, respectively), higher 100 kernel weight (with the means of 28 and 21.67 g for both treatments, respectively), higher yielding ability (with the means of 98.33 and 69.67 g for both treatments, respectively) and consequently more drought tolerant than the other parental line Gm-18. F1 plants had higher performance as for plant height, 100 kernel weight and grain yield per plant and consequently more drought tolerant.
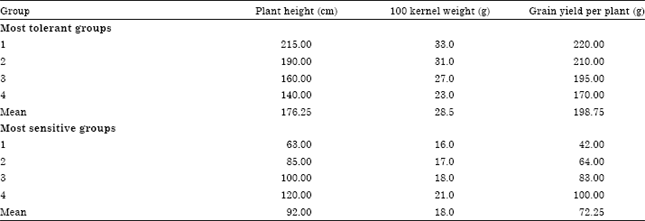
After the two selected maize inbred lines i.e., Sd-63 (tolerant) and Gm-18 (sensitive) were crossed to obtain F1 grains, some of these F1 grains were sown in the field and selfed to obtain F2 grains. Drought experiment was conducted to investigate the response of the F2 segregating population to water stress. F2 plants were classified in a descending order, based on their tolerance to water stress, into groups of 30 F2 individuals in which the most four tolerant and the most four sensitive groups (Table 3) were selected for subsequent molecular analysis. The four most drought tolerant groups had means of 176.25 cm, 28.59 and 198.75 g for plant height, 100 kernel weight and grain yield per plant, respectively while the four most drought sensitive groups had means of 92 cm, 18 and 72.25 g for the above mentioned traits, respectively.
Molecular genetic analysis: The bulked segregant analysis was adopted in this study (Michelmore et al., 1991) to detect markers for drought tolerance in maize. The bulked segregant analysis identifies markers linked to a molecular trait of interest in the segregating F2 population generated from the hybrid between the two contrasting genotypes (tolerant and sensitive in this study).
| Table 2: | Mean performance of two maize inbred lines and their cross for some agronomic traits under normal and drought conditions |
 | |
| Table 3: | Mean performance of the most drought tolerance and the most drought sensitive F2 groups with respect to some agronomic traits under water stress conditions |
 | |
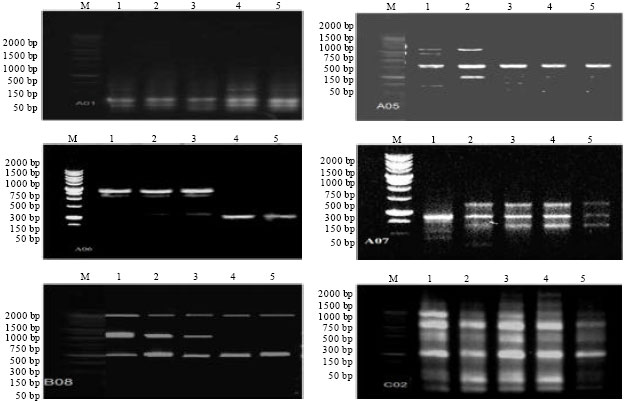 | |
| Fig. 1: | RAPD-PCR profile of (1) tolerant, (5) sensitive maize parents, (3) tolerant F1, (2) the most tolerant F2 and (4) the most sensitive F2 plants (M = marker and RAPD = random amplified polymorphic DNA) |
Two DNA bulks from the most two contrasting F2 groups were used along with their parents and F1 plants to develop RAPD (Random Amplified Polymorphic DNA) markers associated with water stress tolerance. The DNA bulks of F2 of the two extreme groups, for their performance under drought condition, F1 and their parents (Sd-63, tolerant and Gm-18, sensitive) were tested against twenty 10-mer random primers. Data were considered for only six out of twenty primers (Fig. 1), banding pattern for the six primers (A01, A05, A06, A07, B08 and C02) were illustrated in Fig. 1 and scored as present (1) or absent (0) as shown in Table 4. Four out of the six primers (A01, A05, A06 and B08) were developed molecular markers for drought tolerant as shown in Table 4.
| Table 4: | RAPD fragments of six RAPD primers with tolerant (1) and sensitive maize parents (5), tolerant F1 (3), the most tolerant F2 (2) and the most sensitive F2 (4) plants (MW = molecular weight, RAPD = random amplified polymorphic DNA) |
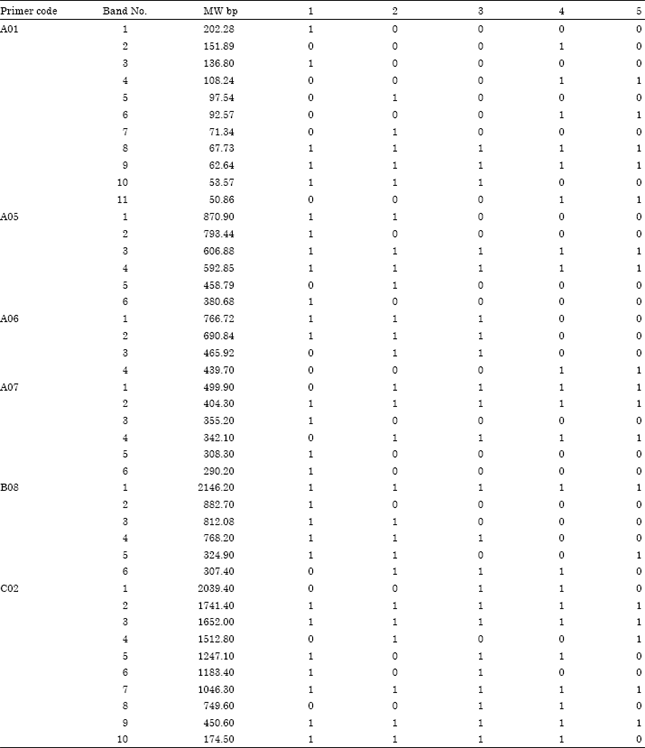 | |
For PCR reaction with the primer A01, two universal bands at Molecular Weights (MW) 97.54 and 71.34 bp characterized the most tolerant F2 group. In the same time, three bands at MW 151.89, 108.24 and 92.57 bp were found in the sensitive parent (Gm-18) and the most sensitive F2 group while were absent in the tolerant groups.
With respect to PCR reaction with the primer A05, two bands at MW 870.90 and 458.79 bp were characterized the tolerant parent (Sd-63) and the most tolerant F2 group while were absent in the sensitive groups.
Regarding PCR reaction with the primer A06, three universal bands at MW 766.72, 690.84 and 465.92 bp were showed to be present for the tolerant groups (the tolerant parent Sd-63, the most tolerant F2 group and the tolerant F1) while were absent in the sensitive groups. In the same time one universal band at MW 439.7 bp was characterized the sensitive parent Gm-18 and the most sensitive F2 while was absent in the tolerant groups.
With respect to PCR reaction with the primer B08, two bands at MW 812.08 and 768.2 bp were showed to be present for the tolerant groups (tolerant parent, F2 and F1) while were absent in the sensitive groups.
Regarding the other two primers (A07 and C02), the data of these primers were not developed any molecular markers for drought tolerant in maize.
The results of the present study are in harmony with the findings of Abdel-Tawab et al. (2002) and Abdel-Bary et al. (2005), they detected some RAPD markers associated with drought and salt tolerance in maize. Rashed et al. (2006) and Younis et al. (2007), they detected some RAPD markers associated with salt tolerance in Sorghum.
CONCLUSION
Results of this study which contain two maize inbred lines, drought tolerant (Sd-63) and drought sensitive (Gm-18), their F1 generation and bulks of the two extreme F2 plant groups (the most tolerant F2 group and the most sensitive F2 group) were tested against six RAPD primers to determine some molecular markers for drought tolerance in maize. This analysis revealed that, four RAPD primers (A01, A05, A06 and B08) out of six developed molecular markers for drought tolerance in maize. These RAPD markers could be considered as reliable molecular markers associated with drought tolerance in maize that can be utilized during breeding programs via marker-assisted selection.
REFERENCES
- Bruce, W.B., G.O. Edmeades and T.C. Barker, 2002. Molecular and physiological approaches to maize improvement for drought tolerance. J. Exp. Bot., 53: 13-25.
CrossRefPubMedDirect Link - Michelmore, R.W., I. Paran and R.V. Kesseli, 1991. Identification of markers linked to disease-resistance genes by bulked segregant analysis: A rapid method to detect markers in specific genomic regions by using segregating populations. Proc. Nat. Acad. Sci. USA., 88: 9828-9832.
PubMedDirect Link - Shiri, M., R.T. Aliyev and R. Choukan, 2010. Water stress effects on combining ability and gene action of yield and genetic properties of drought tolerance indices in maize. Res. J. Environ. Sci., 4: 75-84.
CrossRefDirect Link - De Souza, L.V., G.V. Miranda, J.C.C. Galvao, L.J.M. Guimaraes and I.C. dos Santos, 2009. Combining ability of maize grain yield under different levels of environmental stress. Pesquisa Agropecuaria Brasileira, 40: 1297-1303.
Direct Link - Williams, J.G.K., A.R. Kubelik, K.J. Livak, J.A. Rafalski and S.V. Tingey, 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res., 18: 6531-6535.
CrossRefPubMedDirect Link - Younis, R.A.A., M.F. Ahmed and M.M. El-Menshawy, 2007. Molecular genetic markers associated with salt tolerance in grain sorghum. Arab J. Biotechnol., 10: 249-258.
Direct Link








