Commentary
Mobility of Antibiotic Resistance Through Pharmaceutical Products: A Microbiological Prospective Risk
Department of Microbiology and Immunology, Cairo University, Cairo, Egypt
LiveDNA: 20.8199
Antibiotic resistance has become a globally known threat that may impact not only human health but even life. Dissemination of multi-antimicrobial resistant microbes has become a popular phenomenon that has impacted the public health1. Moreover, there is evidence for the ability of the circular DNA plasmid that is responsible for antimicrobial drug resistance to maintain its functionality even after exposure to unfavorable environmental conditions. Researchers of this study have found that Escherichia coli could uptake this DNA and acquired drug resistance2. Thus, it is surprising to find drug-resistant microbial strains isolated from toilet seats in the International Space Station (ISS) where the environment is confined with limited access and far from the open ecosystem of the planet earth3. Thus, the risk encountered mainly emerges from the resilient nature of antibiotic resistance DNA and the ease of transfer and exchange between micro-organisms. Figure 1 illustrated the exceptional and fast ability of microbial cells to develop drug resistance to the most antimicrobials applied for the cure of infections4,5. An interesting observation showed that the resistance of Pneumococcus to Penicillin was found 22 years after its introduction while Levofloxacin resistance was detected in the same year 1996. The earliest evidence of antibiotic resistance was Staphylococcus to Penicillin5 in 1940. Gram-negative rods were also contributors to the drug resistance such as Shigella to Tetracycline and Enterobacteriaceae to Imipenem and Ceftazidime. Based on centers for disease control and prevention (CDC) records, staphylococcal resistance is one of the most observed resistance against antimicrobial drugs5.
The diversity of bacteria in human, water and environment is of special concern to the microbiological Quality Control (QC) in the pharmaceutical industry not only to detect objectionable micro-organisms but also to investigate the kinetics and mobility of the genetic materials that are responsible to the antibiotic resistance. The swapping process of genetic material between contemporary adjacent bacterial cells may be referred to as horizontal gene transfer6. Various Antibiotic Resistance Genes (ARGs) are integrated on plasmids, transposons or integrons that works as carriers vectors for the transfer of these genes to the other cells of the same species, different species or even another genus7. Three principal mechanisms are responsible for horizontal gene transfer viz transformation, transduction or conjugation. Antibiotic resistance mechanism can occur generally via target modification, efflux, enzyme-catalyzed destruction, immunity and bypass8. In an open ecological framework on earth, it is expected that selective pressure from the exposure to the antimicrobials in nature in a closed loop will be transferred to individuals who will be subjected to drug-resistant bacteria even they have not exposed to these antibiotics before9.
Despite the rules implemented to establish antibiotic treatment protocol as defense lines against the progression of resistance, there are still other routes through which unintentional selection for the most survivors micro-organisms could prevail. The risk of transfer of antimicrobial resistance from food to the final human consumers has been addressed previously by some researchers10,11. Some drugs promote horizontal gene transfer, for example the antiepileptic drug Carbamazepine12. Water and air is also another possible source of the transfer of resistance among other environmental sources that should be investigated13,14. Environmental Monitoring (EM) and water samples in a pharmaceutical facility showed the diversity of micro-organisms that could be found from different sources such as personals, water, surfaces and air15. Even raw ingredients and materials for pharmaceutical product formulation are a possible source for ARGs transfer depending on their microbiological quality and purity16.
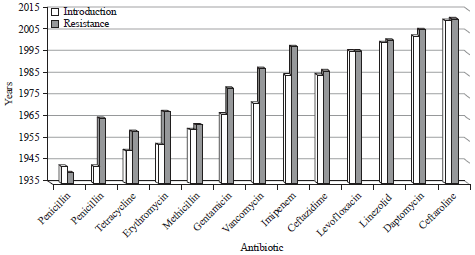 | |
| Fig. 1: | Recorded years of observed microbial resistance for some antimicrobial drugs since their introduction |
| Source: Marty et al.4 and CDC5 | |
| Table 1: | Natural and man-caused selective dynamic spreading of ARGs in nature through main interchangeable sources |
 | |
| Source: Allen et al., 17 | |
Table 1 demonstrated the possible pools of ARGs which are dynamic and mobile between them17.
Although conventional or culture-based techniques are considerably cheap and reproducible, they are known to underestimate the total number of bacteria by up to several orders of magnitude, even with prolonged incubation times and temperature variations. It has long been documented that artificial culture media lead to only a very small fraction 1 in 10000 to 1 in 100 of the total viable bacterial count present in any given sample18. Viable but non-culturable (VBNC) cells have also been shown to be antibiotic tolerant, as well as exhibiting tolerance to heavy metals, environmental stress conditions and ethanol19. Therefore, although significant financial/legal costs linked to culture media-based results, application of selective agents in any culture-based method, including those for pathogens, is likely to lead to a considerable skipping bioburden fraction from the actual number of potentially infective bacteria present. The distinction between “persisters” and VBNC cells appear artificial and while there has been much of work published in recent years on the former, the literature on the latter is voluminous and to date there are no convincing studies indicating they are not the same or at least variants of the same phenomenon20. Interestingly, drug-resistance genes can withstand harsh environmental conditions and reach microbiome of the patient to be absorbed by bacteria as indicated earlier in a previous study2. Thus, current QC tests in pharmaceutical microbiology laboratories under any circumstances are not indicative for the risk associated with antibiotic resistance transfer through pharmaceutical dosage forms, especially if classical official methods were used instead of the new Rapid Microbiological Methods (RMMs) and technologies which are not common in the developing countries particularly.
No reported work relating the role of pharmaceutical manufacturing water, personnel and environment to the mobility of ARG to the final consumers is available till now in our hands. It is encouraged that extensive researches focusing in this dilemma in healthcare industry generally and pharmaceutical sector specifically being performed to access the degree of hazard that could ensue from the kinetics of ARGs from the product to the final consumer who is normally is expected to be in ill-health status and more prone than the normal individuals to the devastating consequences of resistant microbes. Extensive researches are being made to discover new anti-microbial molecules. Nevertheless, there is a great concern over the possibility of balance shifting towards the emergence of the aggressive resistance against antimicrobials in a higher rate rather than susceptibility due to the high adaptability of the microbial cells.