Research Article
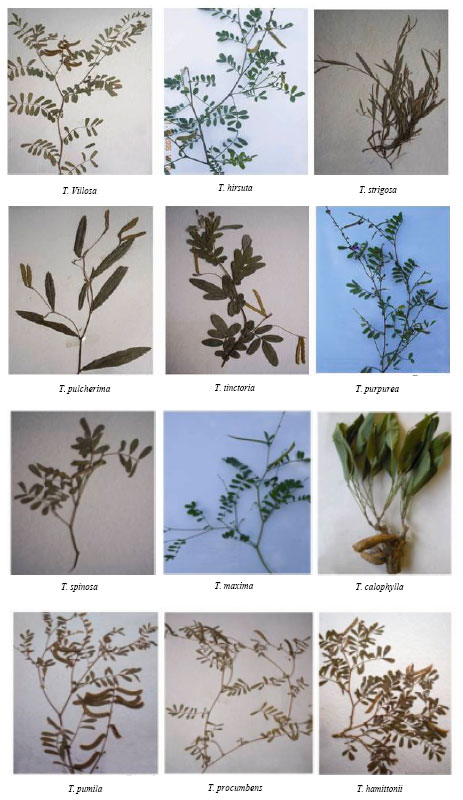
Genetic Relationship Among Tephrosia Species as Revealed by RAPD Analysis
Department of Botany, Andhra University, Visakhapatnam, 530 003 Andhra Pradesh, India
P. Akbar Ali Khan
Department of Pharmaceutical Sciences, Andhra University, Visakhapatnam, 530 003 Andhra Pradesh, India
P. Narasimha Reddy
Department of Pharmaceutical Sciences, Andhra University, Visakhapatnam, 530 003 Andhra Pradesh, India
K. Lakshminarayana
Department of Botany, Andhra University, Visakhapatnam, 530 003 Andhra Pradesh, India
S. Ganapaty
Department of Pharmaceutical Sciences, Andhra University, Visakhapatnam, 530 003 Andhra Pradesh, India











Dayo Reply
Good work....it helps me in my project and seminar...thanks for such a good work..that helps me well.