Research Article
Molecular Modelling Analysis of the Metabolism of Maraviroc
Discipline of Biomedical Science, School of Medical Sciences, Faculty of Medicine, Cumberland Campus, C42, The University of Sydney, Lidcombe, NSW, Australia
Maraviroc (MVC; UK-427,857; 4,4-difluoro-N-{(1S)-3-(exo-3-(3-isopropyl-5-methyl-4H-1,2,4-triazol-4-yl)-8-azabicyclo(3.2.1)oct-8-yl)-1-phenylpropyl}cyclohexanecarboxamide) is a highly selective orally bioavailable chemokine receptor 5 (CCR5) antagonist with potent activity and favourable pharmacological properties against human immunodeficiency virus type 1 (HIV-1) (Dorr et al., 2005; Murga et al., 2006). Since CCR5 is the target of HIV, it is thought that mutation in the gene encoding the receptor would result in immunity to HIV infection. Chemokines are a family of small peptides that act as chemoattractants and activators of specific types of leukocytes. The activated leukocytes can mediate a variety of immune and inflammatory responses through their binding and subsequent activation of specific receptors on the cell surface (Napier et al., 2005).
MVC has been found to show potent antiviral activity against all CCR5-tropic HIV-1 viruses tested, including 43 primary isolates from various clades and diverse geographic origin (Dorr et al., 2005). It has been suggested that significant structural differences between MVC and previously reported CCR5 antagonists underpin the differences in their pharmacological and pharmacokinetic properties (Dorr et al., 2005).MVC is moderately lipophilic and slightly basic in character (pKa = 7.3). It has no detectable in vitro cytotoxicity and is highly selective for CCR5, indicating the potential for an excellent clinical safety profile. MVC is metabolized in humans and experimental animals via multiple pathways to produce a number of metabolites. However, the parent compound is the major circulating and excreted component in all species. UK-408,027 is formed from N-dealkylation at the site adjacent to tropane ring. Methyl oxidation of the triazole ring of UK-408,027 produces MVCM1. Aromatic hydroxylatio at the para position of the phenyl ring of MVC produces UK-437,719.
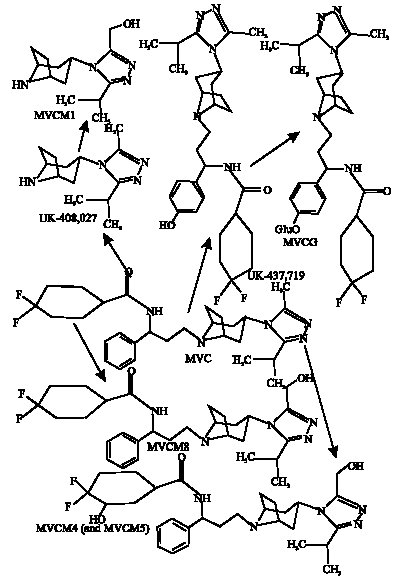 | |
| Fig. 1: | Metabolic pathways for MVC in human, rat, dog and mouse (Walker et al., 2005) (MVCM5, MVCM6 and MVCM7 not shown). |
Oxidation in the difluorocyclohexyl ring produces Met 4 (3HMVC) and Met 5 (2HMVC). Hydroxylation of the azole ring of MVC produces Met 6. UK-437,719 can be glucuroniated to produce corresponding phase 2 metabolite (MVCG). Mono-oxidation of the isopropyl substituent of the azole ring produces two isomers denoted as MVCM6 and MVCM7.Figure 1 summarizes the metabolic pathways for MVC in humans and animals.
In this study, molecular modelling analyses have been carried out using the program Spartan 02 (Spartan, 2002) of MVC and its metabolites, with the aim of providing a better understanding of their toxicity. Molecular modelling analysis is now a mature discipline increasingly used to supply information about structures, relative stabilities and other properties of isolated molecules such as surface charge distribution, surface area, volume, heat of formation and dipole moment (Warren, 2003). For example, it can be shown that whereas the observed heat of formation of propane is -25.0 kcal mol-1, the value calculated using DFT caculation based on 6-31G* basis set is -23.0 kcal mol-1 and the observed C-N bond distance is 0.1358 nm, the value obtained from DFT calculation is 0.1357 nm. Previous studies have shown that xenobiotics and their metabolites which are kinetically labile and abound in electron-deficient regions on the molecular surface tend to induce cellular toxicity due to glutathione depletion and cause DNA damage due to oxidation of nucleobases in DNA (Huq, 2006a, b). The work was carried out in the Discipline of Biomedical Science, School of Medical Sciences, The University of Sydney during March to May 2007.
COMPUTATIONAL METHODS
The geometries of MVC and its metabolites MVCM1, UK-408,027, UK-437,719, MVCM4, MVCM5, MVCM6, MVCM7, MVCM8 and MVCG have been optimised based on molecular mechanics, semi-empirical and DFT calculations, using the molecular modelling program Spartan 02. Molecular mechanics calculations were carried out using MMFF force field. Semi-empirical calculations were carried out using the routine PM3. DFT calculations were carried at B3LYP/6-31G* level. In optimization calculations, a RMS gradient of 0.001 was set as the terminating condition. For the optimised structures, single point calculations were carried out to give heat of formation, enthalpy, entropy, free energy, dipole moment, solvation energy, energies for HOMO and LUMO. The order of calculations: molecular mechanics followed by semi-empirical followed by DFT ensured that the structure was not embedded in a local minimum. To further check whether the global minimum was reached, some calculations were carried out with improvable structures. It was found that when the stated order was followed, structure corresponding to the global minimum or close to that could ultimately be reached in all cases. Although RMS gradient of 0.001 may not be sufficiently low for vibrational analysis, it is believed to be sufficient for calculations associated with electronic energy levels (Huq and Alshehri, 2006).
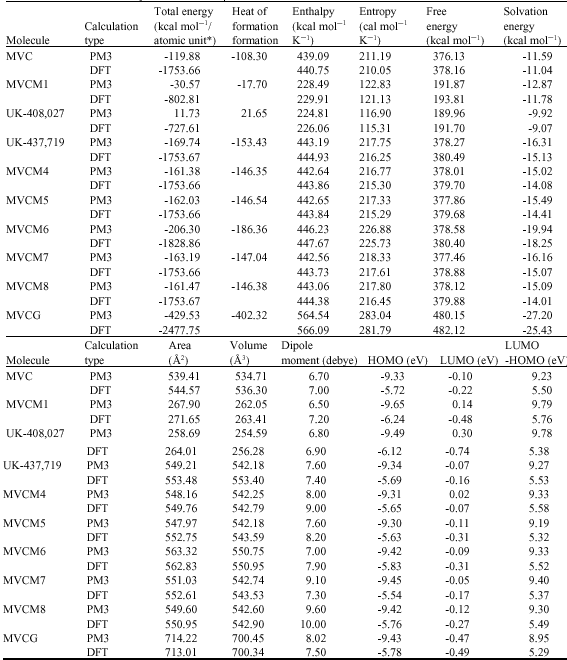
Table 1 gives the total energy, heat of formation as per PM3 calculation, enthalpy, entropy, free energy, surface area, volume, dipole moment and energies of HOMO and LUMO as per both PM3 and DFT calculations for MVC and its metabolites MVCM1, UK-408,027, UK-437,719, MVCM4, MVCM5, MVCM6, MVCM7, MVCM8 and MVCG. Figure 2-11 give the regions of negative electrostatic potential (greyish-white envelopes) in (a), HOMOs (where red indicates HOMOs with high electron density) in (b), LUMOs in (c) and density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) in (d) as applied to optimised structures of MVC and its metabolites MVCM1, UK-408,027, UK-437,719, MVCM4, MVCM5, MVCM6, MVCM7, MVCM8 and MVCG.
The LUMO-HOMO energy differences for MVC and its metabolites MVCM1, UK-408,027, UK-437,719, MVCM4, MVCM5, MVCM6, MVCM7, MVCM8 and MVCG from DFT calculations are found to range from 5.3 to 5.8 eV, indicating that all the compounds would be inert kinetically.
The solvation energies of MVC and its metabolites MVCM1, UK-408,027, UK-437,719, MVCM4, MVCM5, MVCM6, MVCM7, MVCM8 and MVCG obtained from PM3 calculations are found to range from -9.9 to -27.2 kcal mol-1, indicating that the compounds would vary significantly in their solubility in water. When the solvation energy values are compared with the corresponding dipole moments (Table 1), it is found that the order of the solvation energies is not exactly the same as that of the dipole moments. For example, whereas MVCG has the highest solvation energy, it is MVCM8 that has the largest dipole moment.
As noted previously, the results indicate complexity of solution in which such processes as hydrogen bonding and resonance stabilization may be playing key roles (Huq, 2006c).
| Table 1: | Calculated thermodynamic and other parameters of MVC and its metabolites* in atomic units from DFT calculations |
 | |
In the case of MVC, MVCM4, MVCM5 and MVCM7, the electrostatic potential is found to be more negative around the nitrogen centres and fluorine atoms, indicating that the positions may be subject to electrophilic attack. In the case of UK-408,027 and MVCM1, the electrostatic potential is found to be more negative around the various nitrogen centres, indicating that the positions may be subject to electrophilic attack. In the case of MVCM6, MVCG, the electrostatic potential is found to be more negative around the nitrogen and oxygen centres and fluorine atoms, indicating that the positions may be subject to electrophilic attack.
In the case of MVC, MVCM5, the HOMOs with high electron density are found to be close to the non-hydrogen atoms of bicyclorings whereas the LUMOs are found to be close to the non-hydrogen atoms of the phenyl ring. In the case of UK-408,027, MVCM1, MVCM6, MVCM7, UK-437,719, MVCG and MVCM8, the HOMOs with high electron density are found to be close to non-hydrogen atoms of bicyclorings whereas the LUMOs are found to be close to the non-hydrogen atoms of the triazolyl moiety.
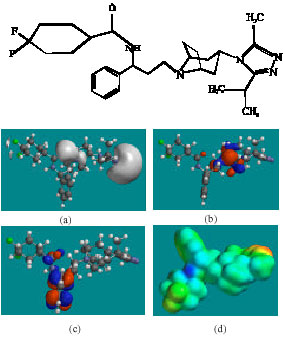 | |
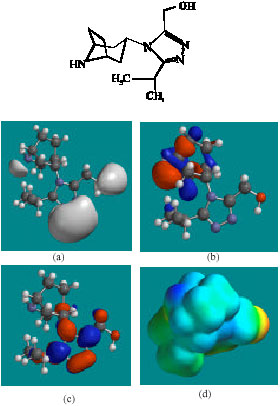
| Fig. 2: | Structure of MVC giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
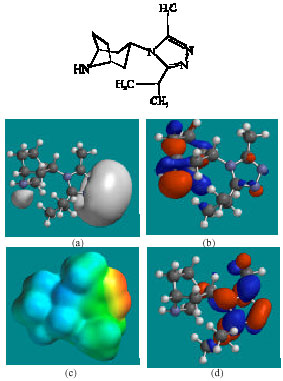 | |
| Fig. 3: | Structure of UK-408,027 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
 | |
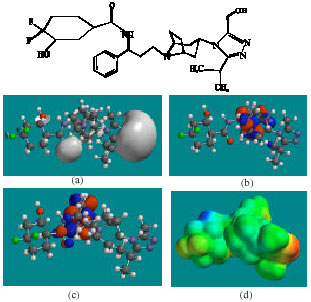
| Fig. 4: | Structure of MVCM1 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral |
 | |
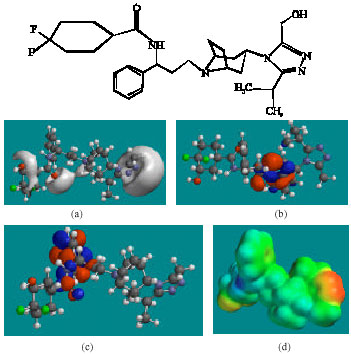
| Fig. 5: | Structure of MVCM4 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
 | |
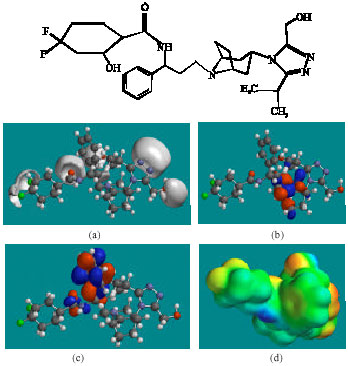
| Fig. 6: | Structure of MVCM5 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
 | |
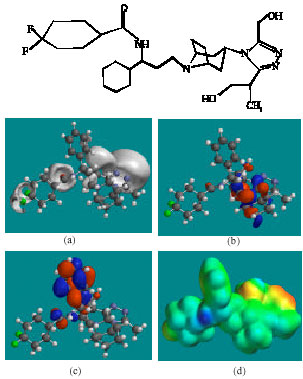
| Fig. 7: | Structure of MVCM6 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
 | |
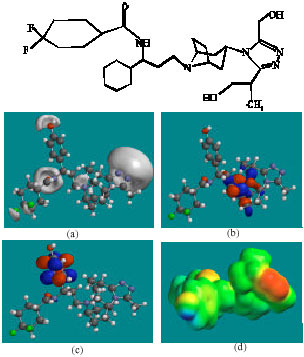
| Fig. 8: | Structure of MVCM7 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
 | |
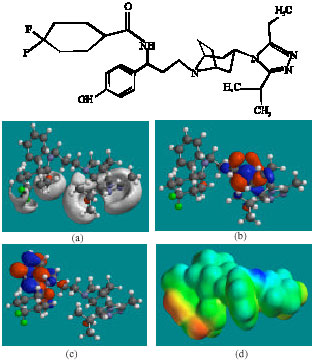
| Fig. 9: | Structure of UK-437,719 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
 | |
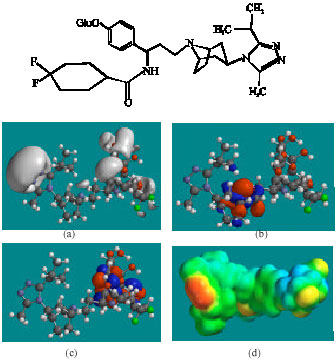
| Fig. 10: | Structure of MVCM8 giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
 | |
| Fig. 11: | Structure of MVCG giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
The disjointness of the HOMOs with high electron density and the LUMOs may be a contributing factor for the kinetic inertness of MVC and UK-408,027. In the case of MVCM4, the HOMOs with high electron density and LUMOs are found to be more closely located.
The molecular surfaces of MVC and all its metabolites are found to abound in neutral (green) regions so that the compounds may be subject to lyophilic attacks. The molecular surfaces of MVC and its metabolites are also found to posssess electron-rich (red and yellow) and electron-deficient (blue) regions so that the compounds may also be subject to electrophilic and nucleophilic attacks. Nucleophilic attacks may be due to glutathione and nucleobases in DNA, resulting into oxidative stress and hence cellular toxicity due to glutathione depletion and DNA damage due to oxidation of nucleobases in DNA. However, the rates of such adverse reactions are expected to be low for MVC and all its metabolites so that none of the compounds may not cause marked cellular toxicity or damage to DNA. Finally, it should be noted that the above results for MVC and its metabolites may explain why MVC has been found to have no detectable in vitro cytotoxicity.
Maraviroc (MVC) is a highly selective orally bioavailable chemokine receptor 5 (CCR5) antagonist with potent activity and favourable pharmacological properties against human immunodeficiency virus type 1 (HIV-1). Molecular modelling analyses based on molecular mechanics, semi-empirical (PM3) and DFT (at B3LYP/6-31G* level) calculations show that MVC and all its metabolites have large LUMO-HOMO energy differences indicating that the compounds would be inert kinetically.This means that although the molecular surfaces of MVC and its metabolites are found to possess some electron-deficient regions so that they may be subject to nucleophilic attacks such as those due to glutathione and nucleobases in DNA, the rates of such adverse reactions would be low unless speed up enzymatically. Thus, MVC therapy may not be associated with severe toxicity.
Fazlul Huq is grateful to the Discipline of Biomedical Science, School of Medical Sciences, The University of Sydney for time release from teaching.