Research Article
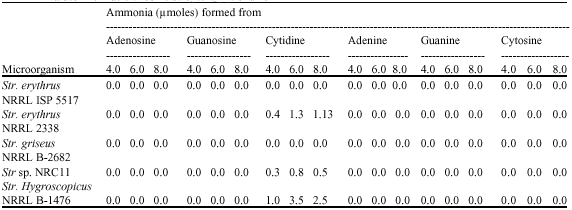
Nucleoside Degradation in Some Streptomyces Strains
Department of Microbial Chemistry, National Research Center, El-Tahrir Street, Dokki, Cairo, Egypt
Latifa A. Mohamed
Department of Microbial Chemistry, National Research Center, El-Tahrir Street, Dokki, Cairo, Egypt