Research Article
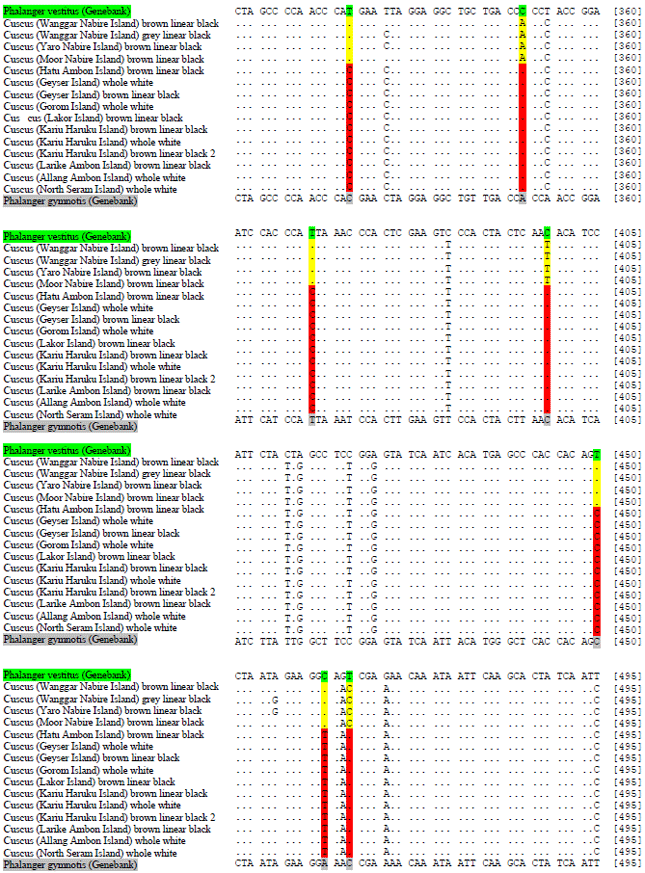
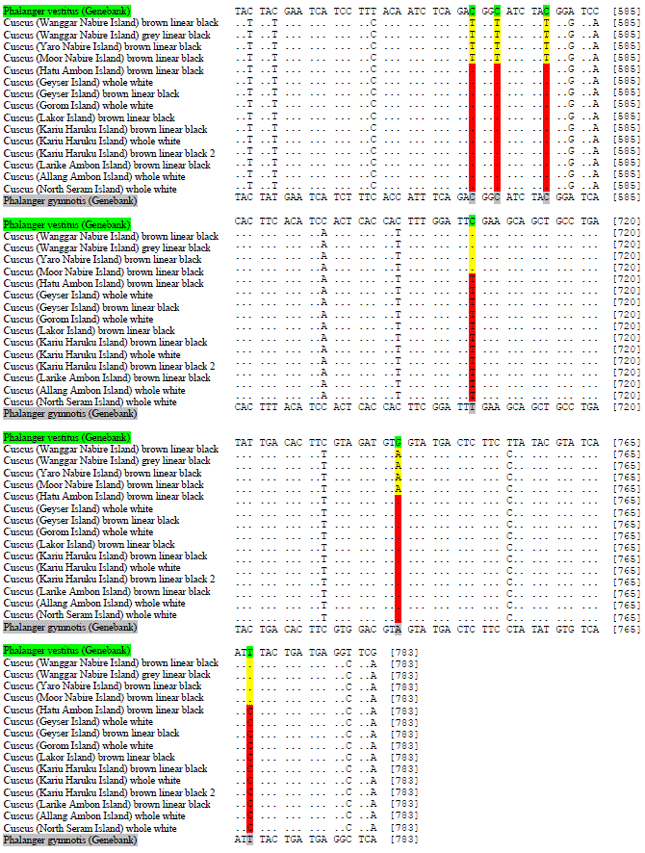
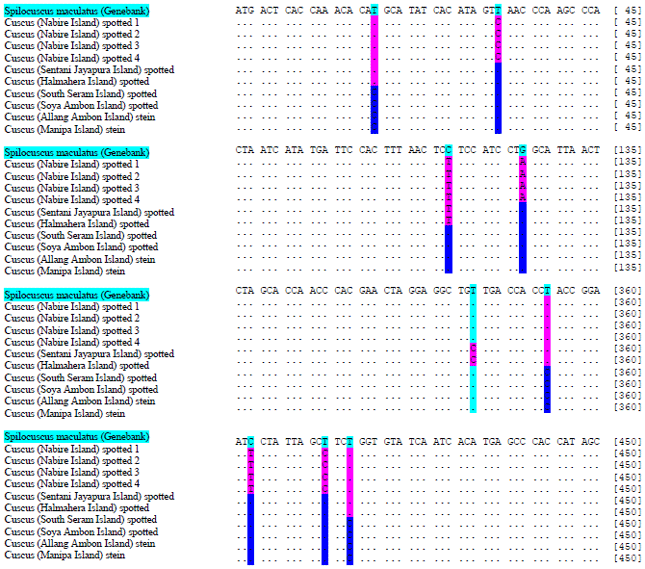
DNA Barcoding of Cuscuses (Marsupialia: Phalangeridae) from Maluku and Papua
Department of Biochemistry, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Yogyakarta, Indonesia
LiveDNA: 62.16743
Niken Satuti Nur Handayani
Department of Genetic, Faculty of Biology, Universitas Gadjah Mada, Yogyakarta, Indonesia
LiveDNA: 62.16385
Hery Wijayanto
Department of Anatomy, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Yogyakarta, Indonesia
Rini Widayanti
Department of Biochemistry, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Yogyakarta, Indonesia
LiveDNA: 62.16531