Research Article
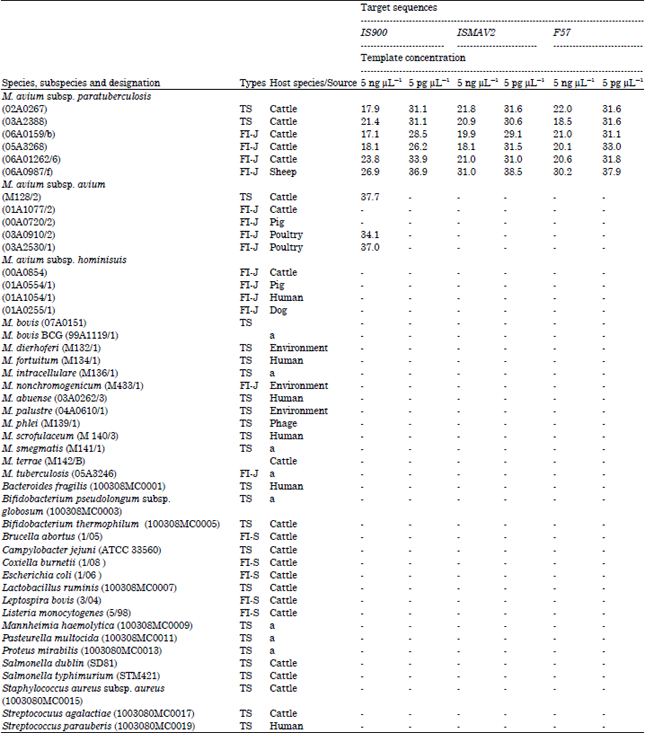
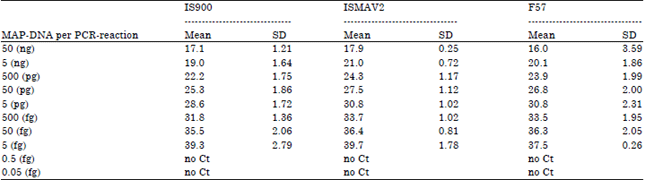
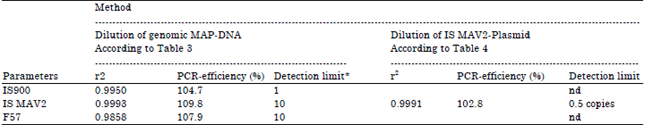
Comparative Evaluation of PCR Assay for Direct Detection of Mycobacterium avium subsp. paratuberculosis in Ruminant
Department of Animal Medicine, Faculty of Veterinary Medicine, Benha University, Moshtohor, 13736, Kaliobia, Egypt
Wolfgang Gaede
State Institute for Consumer Protection of Saxony-Anhalt, Department of Veterinary Medicine, Haferbreiter Weg Str.132-135, D-39576 Stendal, Germany