Research Article
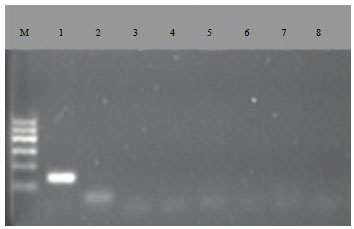
Detection of Mink (Mustela vison) DNA in Meat Products using Polymerase Chain Reaction PCR Assay
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Guangyu Li
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Hanlu Liu
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Wei Zhong
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Haihua Zhang
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Kun Bao
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Chao Xu
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Yahan Yang
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China
Zhuo Wang
Institute of Special Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Jilin, 130112, People`s Republic of China