Research Article
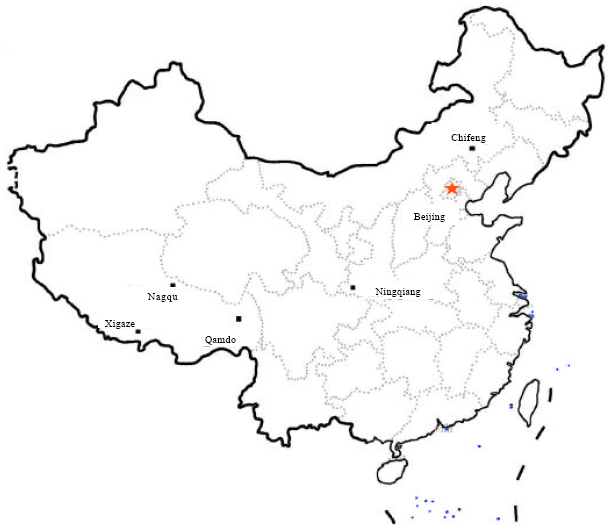
Genetic Diversity of Tibetan Horse and its Relationships with Mongolian Horse and Ningqiang Pony Assessed by Microsatellite Polymorphism
College of Animal Science and Technology, China Agricultural University, Beijing 100193, China
Chunjiang Zhao
College of Animal Science and Technology, China Agricultural University, Beijing 100193, China
Hao Zhang
College of Animal Science and Technology, China Agricultural University, Beijing 100193, China
Guocai Han
College of Animal Science and Technology, China Agricultural University, Beijing 100193, China