Research Article
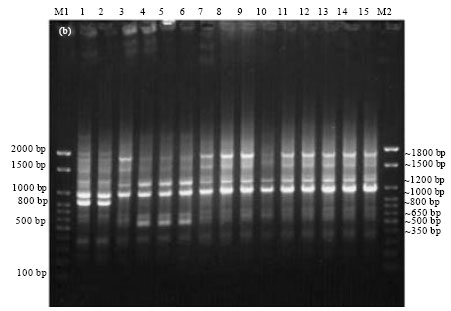
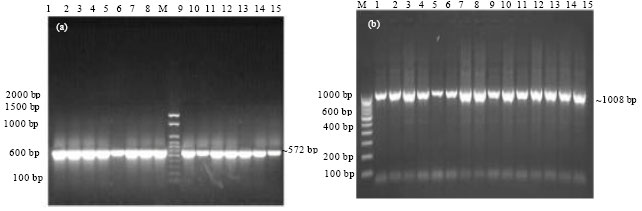
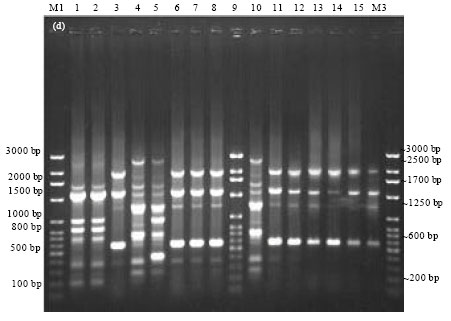
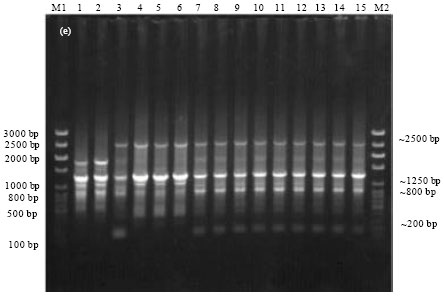
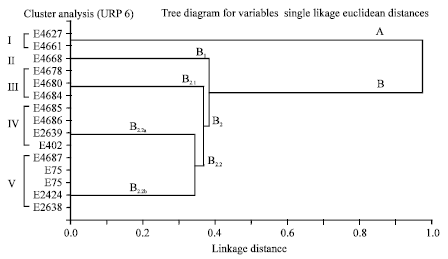
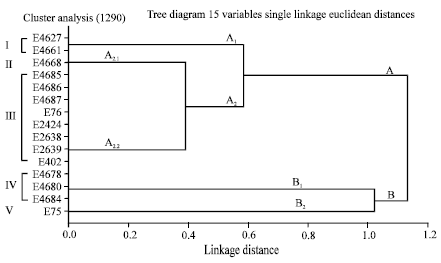
Epidemiological Characterization of Salmonella gallinarum Isolates of Poultry Origin in India, Employing Two PCR Based Typing Methods of RAPD-PCR and PCR-RFLP
Mekelle University College of Veterinary Medicine, P.O. Box 1436, Mekelle, Tigray, Ethiopia
R. Rathore
Centre for Animal Disease Research and Diagnosis (CADRAD), Indian Veterinary Research Institute, Izatnagar, Bareilly (U.P.)-243 122, India
K. Dhama
Avian Diseases Section, Division of Veterinary Pathology, Indian Veterinary Research Institute, Izatnagar (U.P.)-243 122, India
R.K. Agarwal
Division of Veterinary Bacteriology and Mycology, Indian Veterinary Research Institute, Izatnagar (U.P.)-243 122, India