Research Article
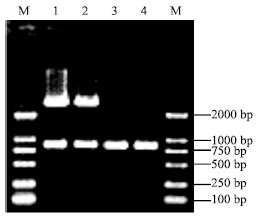
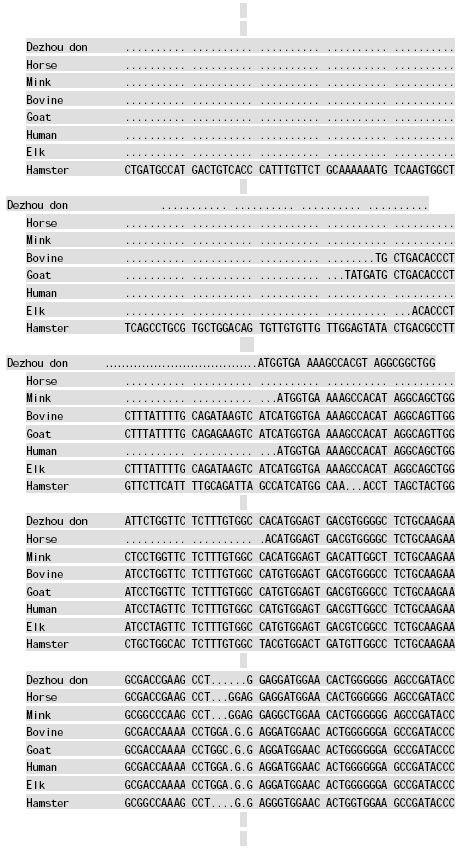
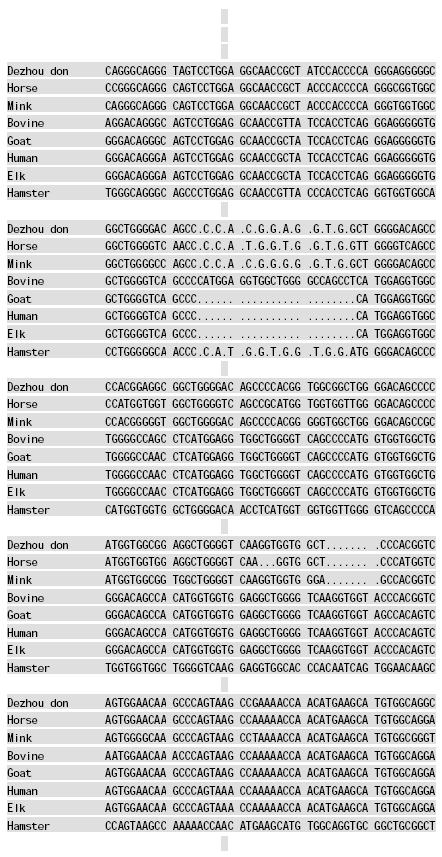
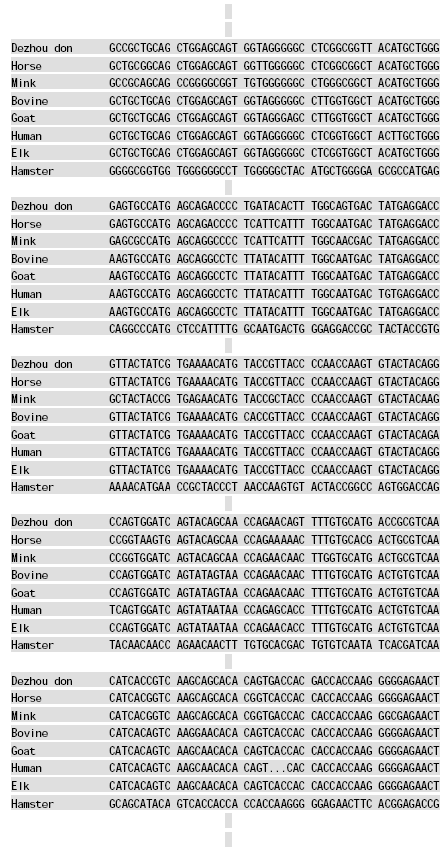
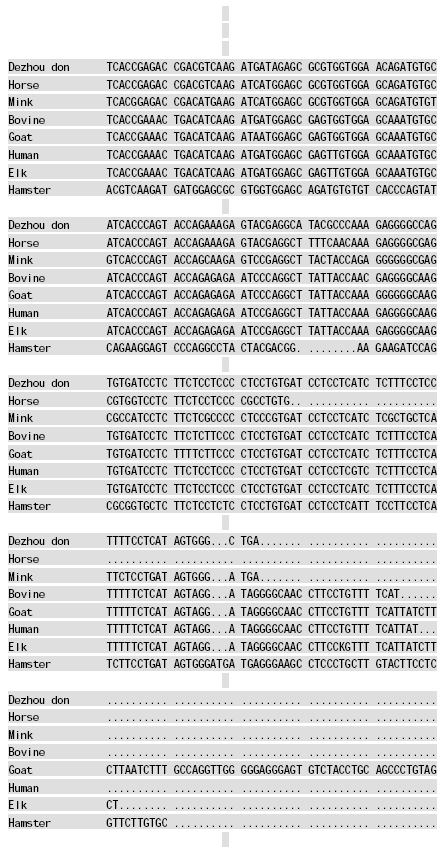
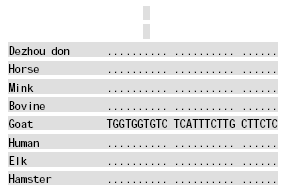
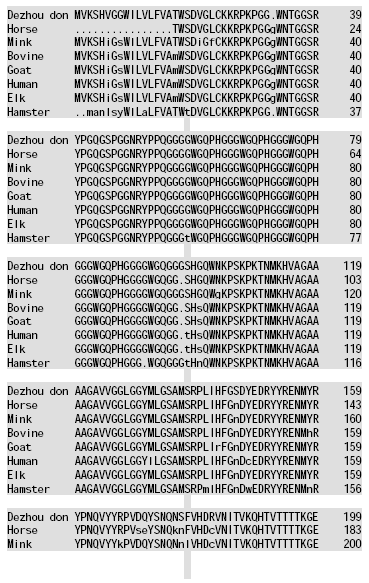
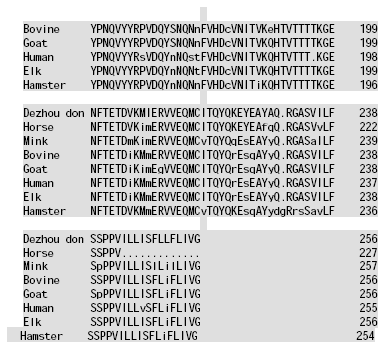
Molecular Cloning and Sequence Analysis of Prion Protein Gene of Dezhou Donkey in China
Laboratory of National Animal TSE, College of Veterinary Medicine, China Agricultural University, Beijing, China
Hongxiang Liu
Departrnent of Histopathology, Addenbrooke’s Hospital, University of Cambridge, Cambridge, UK
Xiangmei Zhou
Laboratory of National Animal TSE, College of Veterinary Medicine, China Agricultural University, Beijing, China
Xiangmei Zhou
Laboratory of National Animal TSE, College of Veterinary Medicine, China Agricultural University, Beijing, China
Deming Zhao
Laboratory of National Animal TSE, College of Veterinary Medicine, China Agricultural University, Beijing, China