Research Article
Using Transposon Mutagenesis to Find an Alternative Resolvase in an Escherichia coli Cells Lacking RuvABC
Department of Genetics, Faculty of Science, The University of Shahrekord, P.O. Box 115, Shahrekord, Iran
Homologous recombination is a fundamental cellular process that promotes the repair of Double Strand Break (DSB) and the rescue of replication forks that have stalled at lesions (McGlynn and Lloyd, 2000, 2002; Michel et al., 2001).
In this process the RuvA, RuvB and RuvC proteins of E. coli encoded by ruvA, ruvB and ruvC genes, respectively are associated with processing of Holliday junction. RuvA and RuvB form a complex that promotes ATP-dependent branch migration of Holliday Junctions (HJ), a process that is important for the extension of heteroduplex DNA. The RuvC protein appears to interact with RuvAB-junction complex to form what has been referred as the RuvABC resolvasome that mediates junction cleavage (West, 1997).
In repair of stalled fork, it was suggested that rewinding of the parental strands and annealing of the nascent strands drive the fork back from the lesion and generate a structure resembling a HJ from a fork. The resolution of this structure by RuvABC provides double strand break that promotes recombination. Thus, RuvABC protein is required for initiation and termination of recombination in this fork repair pathway (McGlynn and Lloyd, 2000; Michel et al., 2001).
Strains lacking RuvA, B or C are unable to process junctions and are equally sensitive to DNA damaging agents such as UV light and mitomycin C (MC) (Mahdi et al., 1996). Due to the importance of these proteins, genes encoding of RuvA, RuvB and RuvC are present in many organisms. In microorganisms having RuvC, including E. coli, some have another resolvase, namely RusA. RusA is a HJ resolvase encoded by rusA gene of cryptic lambdoid prophage DLP12 of E. coli K-12. RusA is normally inactive and deletion mutations showed no obvious effect on recombination or sensitivity to UV light (Mahdi et al., 1996). However, it can be activated following the insertion of IS2 or IS10 containing promoter elements upstream of rusA to activate transcription (Mahdi et al., 1996). Meanwhile, in some microorganisms except E. coli, RusA is the only resolvase that has been already known, such as Aquifex aeolicus (Sharples et al., 1999).
Holliday junctions can also be processed in E. coli by RecG and RusA proteins in the absence of RuvABC. RecG is a DNA helicase that like RuvAB can drive branch migration of HJ and other branched structure (Lloyd and Sharples, 1993). It was shown that it can also form a HJ from a replication fork (McGlynn and Lloyd, 2000; Singleton et al., 2001). Then, RusA cleaves the point of crossover (Sharples et al., 1994; Chan et al., 1998).
Moreover, Mus81 is an endonuclease in yeast and mammals that like RuvC resolves Holliday junction (Osman and Whitby, 2007). The ability of RusA to suppress the UV and MMS sensitivities of a mus81 mutant is consistent with the involvement of Mus81 in repair of stalled forks (Osman and Whitby, 2007).
On the other hand, there are some microorganisms that do not have either RuvC or RusA, including Mycoplasma sp. (Sharples et al., 1999). It suggests either they may have an unknown resolvase or they may use a repair pathway that is not dependent on recombination. Examples of these repair pathways have been presented by Trautinger et al. (2005) and Heller and Marians (2006). The aim of this study was to find an unknown resolvase that promote survival in UV irradiated cells lacking RuvABC.
| Table 1: | E. coli strains used in this study |
 | |
| All strains are MG1655 derivatives. R, S and WR are abbreviation of resistance, sensitive and weak resistance, respectively | |
Bacterial strain: This study was conducted couple of months ago. Escherichia coli K-12 strains used in this study are shown in Table 1. λNK1327 carries Tn10kan plac as described previously by Kleckner et al. (1991). P1vir is a virulent mutant of P1. IPTG (isopropyl-β-D-thiogalactopyranosid) was used in LB broth at 0.022 mg mL-1 to induce transcription from the lac promoter.
Media and general methods: LB broth and agar were used for bacterial culture. Media recipes and procedures for strain construction by P1vir-mediated transduction, testing sensitivity to MC and measuring survival of UV irradiated cells have been cited previously by Jaktaji and Lloyd (2003). UV survival values are means of three or four independent experiments.
Transposon mutagenesis: Tn10kan insertions were generated by infection of strain N4237 with λNK1327 and selection for kanamycin resistant clones at 42°C as described previously. Colonies of resistant clones were regrown in regular arrays and tested for sensitivity to MC and UV light using plate assays as described previously by Jaktaji and Lloyd (2003).
PCR amplication and DNA sequencing: Chromosomal DNA was extracted as cited previously by Jaktaji and Lloyd (2003). Location of insertion upstream of rusA gene was identified by sequencing PCR products amplified from chromosomal DNA using rusA specific primer 5�-TGGCGAAGAAGCTTTGCCATATTA-3� and IS10 specific primer 5�-CACCTATGTGTAGAACAGTATA-3�.
Strain N4237 was mutagenised using a Tn10kan element as described above. kmR clones showing evidence of increased growth on MC agar plates irradiated with UV were purified and further tested. Using this screen, some 10000 KmR derivatives of N4237 were tested.
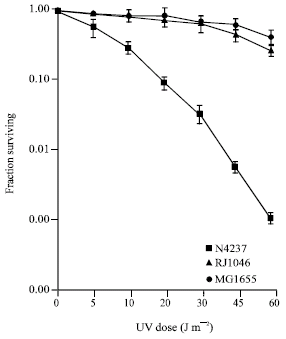 | |
| Fig. 1: | The effect of insertion mutation on UV sensitivity |
The RJ1046 clone was retained for study because it had increased resistance to both UV light (Fig. 1) and MC (data not shown) compared with the N4237 parent and MG1655 wild type strains. Since, the Tn10kan element used carries the plac promoter, it is possible that it activated the expression of a poorly transcribed gene that could promote repair (Kleckner et al., 1991).
It was shown that purE has 50% linkage to rusA (Mandal et al., 1993). To see if RJ1046 carries an insertion that activates rusA, linkage of the Tn10kan insertion to purE85::Tn10 in the ruvA63 strain N3598 was examined. P1 grown on RJ1046 was used to transduce strain N3598 to pur+. These transductants (100) were screened for sensitivity to Km and Tc using plate assays. The inheritance of the purE+ donor marker is associated with loss of Tn10. All pur+ transductants proved MCR. Also 47 out of 100 proved KmR. These results indicated that the Tn10kan insertion in RJ1046 was linked to rusA. To confirm above results PCR amplification of the fusion and DNA sequencing were conducted as described above using strain RJ1054. The data drawn from PCR analysis showed an insertion is in orf96 upstream of rusA (Fig. 2). As mentioned above the Tn10kan element carries the plac promoter. To see if induction of this promoter to over express rusA could further improve survival, the RJ1046 was tested for sensitivity to UV light in the presence and absence of IPTG (an inducer of plac promoter). The results showed that addition of IPTG had minor effect on UV survival of RJ1046 (Table 2). This indicates that the presence of the insertion is sufficient to allow enough expression of rusA to promote repair.
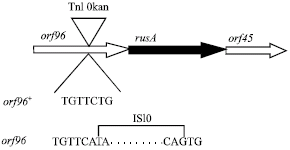 | |
| Fig. 2: | Location of Tn10kan insertion in the orf96 gene |
| Table 2: | The effect of expression (IPTG minus) and over expression (IPTG plus) of rusA on UV survival |
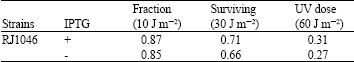 | |
| The data are means of three independent experiments. Errors were less than 10% of the mean values | |
RuvC resolvase in conjunction with RuvAB branch migration motor produces RuvABC resolvasome (West, 1997). RuvABC involves in breaking the stalled fork and providing substrate for initiation of recombination. This protein also participates in processing of recombination intermediate to convert it to fork structure (McGlynn and Lloyd, 2000, 2002; Michel et al., 2001).
Homologs of RuvC protein have been found in many organisms from phages to human. Constantinou et al. (2001) found a combined branch migration and resolution activity in mammalian cell extracts. The human and yeast Mms4-Mus81 endonuclease (homolog of RuvC) arise when DNA replication is blocked by damage or by nucleotide depletion (Constantinou et al., 2001; Chen et al., 2001; Osman and Whitby, 2007).
In an attempt to find a substitute for RuvABC protein in E. coli cells lacking RuvABC, one UV and MC resistant clone, RJ1046 was obtained. By transductional mapping the location of Tn10kan was found which is near rusA gene. Moreover, by PCR sequencing the exact location of Tn10kan insertion was determined. It is in orf96 gene, upstream of rusA gene that activates the expression of it. RusA is the Holliday junction-specific endonuclease (Sharples et al., 1994; Chan et al., 1998) that needs a branch migration protein, RecG to act (Lloyd and Sharples, 1993). However, unlike RuvC, it does not need to bind with a DNA branch migration protein like RecG or RuvAB to be activated (Rafferty et al., 2003).
This study and earlier study findings by Mahdi et al. (1996) that showed deletion or over expression of rusA in the presence of IPTG has no obvious effect on recombination and sensitivity to UV light suggests that although RusA can be activated in the absence of RuvABC, it is not vital for cell survival. Moreover, McGlynn and Lloyd (2000) found that activation of RusA in ruv mutant lacking recB has severe effect on sensitivity to UV light.
Finally, this study could not find an unknown resolvase in E. coli cells. This suggests that in the absence of RuvABC, E. coli cells may use an alternative non-recombinational repair pathway to rescue stalled fork. This might be a case for microorganisms that do not have both RuvC and RusA. A model of this non-recombinational repair pathway type was presented by Lloyd and Sharples (1993) and Trautinger et al. (2005). More recently, it was suggested that even in the presence of RuvABC, cells could have a strategy to prevent unnecessary recombination during replication fork repair (Mahdi et al., 2006).
Strains used in this study were Prof. R. G. Lloyd’s laboratory strains. The author was supported by University of Shahrekord.