ABSTRACT
Extensive genetic variation exists within peanut (Arachis hypogaea L.) germplasm. For a small-scale screening of resistance to tomato spotted wilt virus (TSWV) and flavonoid content, twenty-four accessions were selected from peanut germplasm in the US collection and planted in the greenhouse. Plant response to TSWV was observed and recorded. Leaf tissues were collected and tested by an Enzyme-Linked Immunosorbent Assay (ELISA) for TSWV. Harvested seeds from mature plants were used for quantification of flavonoid content by High Performance Liquid Chromatography (HPLC) analysis. Seed-coat colors were also recorded. One highly resistant and eight moderately tolerant accessions to TSWV were identified within A. hypogaea and confirmed by ELISA. Peanut seeds contained trace amounts of myricetin and genistein and a low amount of daidzein and kaempferol, whereas some accessions contained a high amount of quercetin. Intriguingly, all accessions with a high amount of quercetin had a white seed-coat color. There was no clear association between resistance to TSWV and amount of quercetin observed in the accessions evaluated in this experiment.
PDF Abstract XML References
How to cite this article
DOI: 10.3923/ppj.2007.219.226
URL: https://scialert.net/abstract/?doi=ppj.2007.219.226
INTRODUCTION
Cultivated peanut (Arachis hypogaea L., 2n = 4x = 40) is a tetraploid species and one of the five most important oilseed crops in the world. The United States is one of the major peanut producers and exporters (Carley and Fletcher, 1995). Peanut seeds contain approximately 50% high quality oil and 25% protein. Although peanuts are traditionally used for oil production, they can be processed (i.e., boiled, roasted, salted, fried or fermented) into various forms of foods such as peanut butter, nuts, candy, snacks and confections for human consumption. Due to their high protein content, peanuts were also used as protein supplements in cereals, other legumes, or root-based food products to improve food quality or alleviate malnutrition caused by protein-deficiency in the world (Singh and Singh, 1991).
Flavonoids (including daidzein, genistein, kaempferol, myricetin and quercetin) as secondary metabolites have beneficial effects on human health due to their antioxidant, antiestrogenic and antiproliferative activities. Recently, it was found that peanut seeds contain a negligible amount of myricetin, trace amounts of daidzein, genistein and kaempferol and a higher amount of quercetin than other legume seeds (Wang et al., 2007). Within the same genotype, peanut seedlings possess a much higher concentration of isoflavones and resveratrol than seeds (Wang et al., 2005; Kirakosyan et al., 2007). Peanut seedlings have been used as vegetables in Asia. The high concentration of isoflavones and resveratrol in the peanut seedlings should add more value for recommending peanut seedlings as nutritious and functional vegetables.
Tomato spotted wilt virus (genus Tospovirus, family Bunyaviride, TSWV) has a wide range of hosts and infects over 800 plant species, including peanut (German et al., 1992; Jones, 2004; Hurt et al., 2005). It is a serious threat to peanut production and causes a significant yield loss worldwide. The first spotted wilt on peanuts in the United States was observed in Texas in 1971. Since then, it has become one of the most serious diseases of peanuts. Annual losses on peanuts due to TSWV were estimated to be about $40-100 million in Georgia alone (Jain et al., 1998; Culbreath et al., 2003). Since TSWV in peanuts is transmitted by vector thrips, it is difficult to control. There is no single measure which can be used to control TSWV effectively. An integrated approach (including use of resistant cultivars, application
of chemicals and practice of cultivation) has been developed to manage TSWV in peanuts (Todd et al., 1998; Culbreath et al., 2003).
Germplasm accessions highly resistant to TSWV have been identified by screening wild forms from 20 diploid species (Lyerly et al., 2002). Breeding lines highly resistant to TSWV have been developed by crossing and screening interspecific progenies (Holbrook et al., 2003). However, germplasm accessions within A. hypogaea were not well screened and high levels of resistance to TSWV germplasm have not been found (Lyerly et al., 2002). For improvement of TSWV resistance, genetic resources from A. hypogaea (tetraploid) would be easier to use than from diploid species in breeding programs. Therefore, germplasm from A. hypogaea needs to be screened for resistance to TSWV. Variation in flavonoid content was also observed in peanuts (Wang et al., 2007). Furthermore, some flavonoids were demonstrated to have antiviral activities (Clark et al., 1998; Li et al., 2000, 2002; Jassim and Naji, 2003). Quercetin is the most abundant of flavonoid molecules, existing in roots, leaves, stem, flowers and seeds (Harris et al., 2007). Quercetin showed strong antiviral activities for herpesviruses (HSV) and adenoviruses on mammalian cells (ADV) (Chiang et al., 2003; Ozcelik et al., 2006) and strong inhibitory activities for aphid reproduction on cowpea plants (Lattanzio et al., 2000). Since some peanut accessions contain a high amount of quercetin, peanut plants may be able to use quercetin for anti-TSWV activities. The objectives of this study were to identify peanut accessions highly or moderately tolerant to TSWV, to identify peanut accessions with a high content of flavonoids and to determine whether there was an association between a high content of quercetin and resistance to TSWV.
MATERIALS AND METHODS
Twenty-four peanut accessions within A. hypogaea L. species were selected from the US germplasm collection. Their accession number, origin or collection site and identifier are shown in Table 1. Six peanut seeds from each accession were planted in trays in the Griffin greenhouse in 2005. Two treatments were arranged in two different bays. Treatment I was in bay one in which had no thrips or thrip hosts. Therefore, plants in the bay one will have no TSWV infection. Treatment II was in bay two in which had thrips or thrip hosts. Since the thrips contained TSWV, the plants in the bay two will have TSWV infection. When the TSWV symptoms were fully developed, four plants from each accession were scored separately by two scientists for response to TSWV. Based on the severity of plant stunting, the percentage of leaf curling and discoloration, the individual plant was ranked from 1 to 10 (1 = highly resistant, no plant stunting, leaf curling and discoloration; 10 = highly susceptible, severe plant stunting, leaf curling and discoloration).
Fresh leaf tissues from each scored plant were collected and used for TSWV detection by an alkaline phosphatase-based double antibody sandwich enzyme-linked immunosorbent assay (DAS-ELISA) using matching conjugate to the purified antibody used as plate coat with trituration of the leaf samples done employing autoclaved mortars and pestles in leaf buffer for peanut after the method described by Pinnow et al. (1990).
| Table 1: | Selected peanut accessions |
 | |
Finished plates were read at 405 nm on a molecular devices E-max plate reader.
A standard curve for daidzein, genistein, kaempferol, myricetin and quercetin was established on HPLC using standards purchased from Sigma (St. Louis, MO). Seeds were harvested from each individually scored plant and seed-coat colors were observed and recorded. Harvested peanut seeds were used for quantification of flavonoids by HPLC analysis following the method described by Wang et al. (2007). Seeds were ground into a fine powder and used for extraction of flavonoids in 80% methanol containing 1.2 M hydrochloride acid by hydrolysis at 80°C. Supernatant containing flavonoids was filtered through a syringe with a 0.2 μm filter prior to injection into an HPLC system. Flavonoid separation was performed on the Agilent 1100 series HPLC system using a C18 column (150x4.6 mm i.d., 5 μm) with a guard column (12.5x4.6 mm i.d., 5 μm) in a mobile phase. The mobile phase consisted of A: Water (pH 2.5 with formic acid) and B: Acetonitrile (GFS Chemicals, Powell, OH). A gradient elution was carried out at a flow rate of 2.0 mL min-1 with a column temperature of 40°C. The gradient profile was programmed at 20% B from 0 to 2 min, 20-30% B from 2 to 20 min, 100% B from 22 to 24 min, 20% B from 24 to 26 min, then holding at 20% B for 2 min. A 5 min post time was allowed for a system equilibration prior to each sequential injection. The supernatant from the same extraction was injected twice. A software package from Agilent Technologies (ChemStation for LC 3D, reversion A. 10.01) was used for quantification of each flavonoid based on the peak area obtained from HPLC. Each accession was extracted at least three times. The average value from three extractions was used for calculation of flavonoid content.
RESULTS AND DISCUSSION
Plant response to TSWV and confirmation by ELISA: In treatment I, plants were healthy (no TSWV symptom observed) and seeds were harvested from all except two plants (94 plants). In treatment 2, most of the plants showed TSWV symptoms (Fig. 1) and seeds could only be harvested from 45 plants due to either high spotted wilt or only vegetative growth of plants (Table 2). Significant variation on the response to TSWV was observed among different accessions, ranging from highly susceptible to highly resistant (Fig. 1, Table 2). ELISA detection was conducted on leaf tissues because sufficient amounts of tissues could be easily obtained. The observational scores were generally consistent with the value of ELISA detection (the value of ELISA detection >0.1 was considered a positive infection with TSWV). For example, three of the four plants within PI 493582 were scored positive (scored as 5, 7 and 9, respectively); confirmed by ELISA detection (valued as 1.612, 0.618 and 2.824, respectively) and therefore, the accession was rated as highly Susceptible (S). Two of the four plants within PI 262077 were scored positive (scored as 7 and 8); confirmed by ELISA detection (valued as 1.085 and 3.305) and therefore, the accession was rated as Moderately Susceptible (MS). One of the four plants within PI 602506 was scored as TSWV positive (scored as 5); confirmed by ELISA detection (valued as 3.103) and thus, the accession was rated to be Moderately Tolerant (MT). All four plants within PI 509483 were scored as TSWV negative (scored as 1, 1, 1 and 1); confirmed by ELISA-detection (valued as 0.068, 0.043, 0.041 and 0.019, respectively) and thus, the accession was rated as highly Resistant (R). Overall, eight accessions were rated as Susceptible (S), seven accessions as Moderately Susceptible (MS), eight accessions as Moderately Tolerant (MT) and only one accession was rated as highly resistant (R, Table 2).
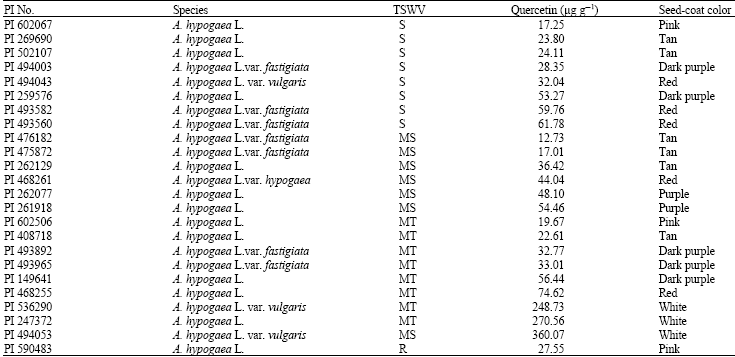
Flavonoid content and seed-coat color: Five flavonoids were compared to commercial standards (in a hydrolyzed form), quantified from harvested seeds by HPLC analysis and expressed in μg g-1 (FW). Peanut seeds contained trace amounts of genistein and myricetin and a low amount of daidzein and kaempferol whereas some accessions contained a high amount of quercetin (Fig. 2). There was significant variation on quercetin content in peanut seeds and peanut seed-coat color. Peanut seed-coat colors varied from white to dark purple (Table 3). PI 247372 contained a higher concentration of quercetin (46.58 ng μL-1) than PI 262077 (11.66 ng μL-1) (Fig. 3). Variation on the quercetin content among peanut accessions tested was from 17.25 to 360.07 μg g-1 (Table 3). Three accessions (PI 536290, PI 247372 and PI 494053) contained much higher amounts of quercetin (248.73, 270.56 and 360.07 μg g-1) than the other 21 accessions. Intriguingly, all three accessions with a high amount of quercetin had a white seed-coat color. This result was consistent with the result from our previous study. PI 247372 (used in this study) was the only accession which had a white seed-coat color in our previous study. It contained a much higher amount of quercetin than the other peanut accessions and the other legume accessions.
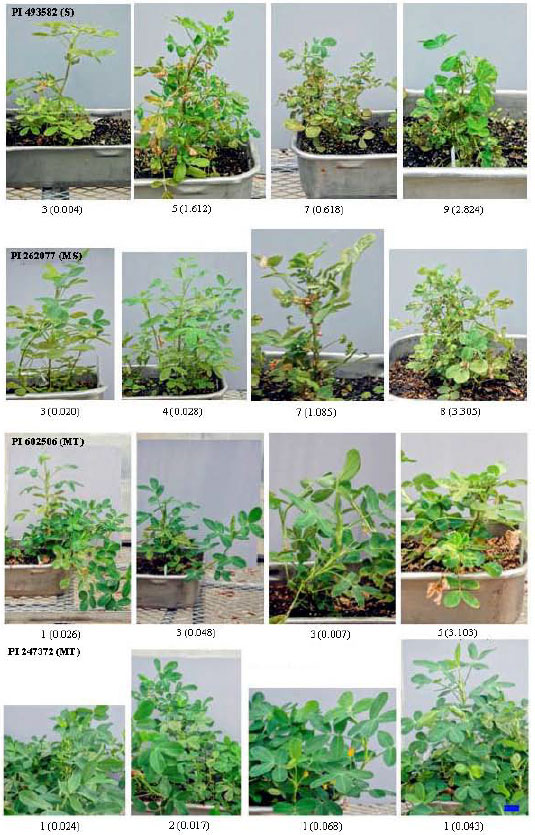 | |
| Fig. 1: | Plant symptoms, observation score and ELISA-detection value for TSWV from different plants and accessions. |
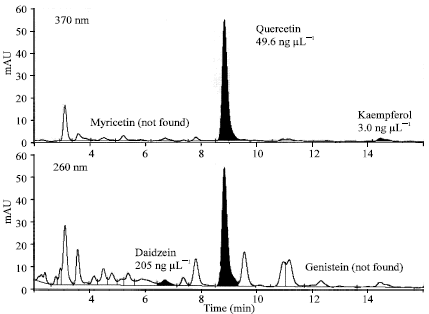 | |
| Fig. 2: | Chromatograms of different flavonoids from HPLC analysis. PI 247372 is used as an example. The upper panel shows a chromatogram at 370 nm on which myricetin is not identified whereas quercetin and kaempferol are identified. The lower panel shows a chromatogram at 260 nm on which genistein is not identified whereas daidzein and quercetin are identified |
 | |
| Fig. 3: | A comparison of quercetin concentrations in the seeds from two peanut accessions on chromatograms generated by HPLC analysis. The upper panel shows the quercetin concentration from the seeds of PI 262077 and the lower panel shows the quercetin concentration from the seeds of PI 247372 |
Since only one peanut accession had a white seed-coat color in our previous study, the conclusion could not be drawn that white seed-coat color was associated with a high content of quercetin in peanuts (Wang et al., 2007). Based on the results from the previous and present studies, a white seed-coat color appears to be associated with a high amount of quercetin in peanut seeds.
| Table 2: | Plant response to TSWV from observational-score in the greenhouse and ELISA-detection |
 | |
| Observation-score from 1 to 10 (1 = Highly resistant; 10 = Highly susceptible); Numbers in the brackets are the values from ELISA detection (Neg = Negative; Pos = Positive). *: Plants did not produce seeds; S: Susceptible, T: Tolerant, R: Resistant and M: Moderate | |
| Table 3: | Quercetin content, seed-coat color and resistance to TSWV of selected peanut accessions |
 | |
To confirm this association, further investigations are warranted. For instance, several white and dark seed-coat color accessions are to be selected from the US collection and grown in the same environment. Seeds collected from the same environment are to be used for quantification of quercetin to determine whether there is a clear difference on the quercetin content between white seed-coat accessions and dark seed-coat accessions.
Amount of quercetin and resistance to TSWV: Although extensive variation was observed on both amount of quercetin in peanut seeds and resistance to TSWV, there was no clear association between resistance to TSWV and amount of quercetin observed in the accessions evaluated in this experiment. For example, PI 590483 was highly resistant to TSWV but it contained a low amount of quercetin (27.55 μg g-1). PI 494053 contained a very high amount of quercetin (360.07 μg g-1) but it was moderately susceptible to TSWV. However, it was observed that PI 536290 and PI 247372 contained a high amount of quercetin (248.73 and 270.56 μg g-1) and they showed moderately tolerant to TSWV. From our experiment, we had not harvested sufficient amount of seeds from the TSWV-challenged plants (treatment 2) to perform HPLC analysis for quantification of quercetin in the seeds. Our greenhouse conditions were not optimum for peanut seed production plus the added stress of TSWV infection. This may explain why we failed to produce enough seeds. Leaf tissues could be used for quantification of quercetin but collection of leaf tissues from plants may affect peanut seed production. That was why leaf tissues were not used for the quantification of quercetin in this experiment. Strong antiviral (herpesviruses and adenoviruses) activities from quercetin were observed in other species (Chiang et al., 2003; Ozcelik et al., 2006). There was also an association observed between total phytoalexin (including resveratrol) production and published genotype resistance to major diseases (including TSWV) (Sobolev et al., 2007). One possible explanation would be peanut seeds probably do not contain sufficient amount of quercetin for exhibiting antiviral activities. TSWV infection may stimulate plants to synthesize more quercetin. To investigate the association between amount of quercetin and resistance to TSWV in the future, peanut seeds need to be planted in the field; plants need to be challenged by TSWV and leaves infected with TSWV and seeds harvested from TSWV-infected plants need to be collected and used for quantification of quercetin content by HPLC analysis.
ACKNOWLEDGMENTS
The authors gratefully thank Mr. Brandon Tonnis for his excellent assistance on HPLC analysis, Ms. Lee Ann Chalkley and Tiffany Fields for providing peanut seeds and Drs. Noelle Barkley, Graves Gillaspie and James Buck for reviewing the manuscript.
REFERENCES
- Jones, R.A.C., 2004. Using epidemiological information to develop effective integrated virus disease management strategies. Virus Res., 100: 5-30.
CrossRefDirect Link - Kirakosyan, A., P.B. Kaufman, J.A. Duke, E.M. Seymour, S. Warber and S.F. Bolling, 2007. Production of isoflavones in seeds and seedlings of different peanut genotypes. Crop Sci., 47: 717-719.
Direct Link - Chiang, L.C., W. Chiang, M.C. Liu and C.C. Liu, 2003. In vitro antiviral activities of Caesalpinia pulcherrima and its related flavonoids. J. Antimicrob. Chemother., 50: 194-198.
CrossRefDirect Link - Lattanzio, V., S. Arpaia, A. Cardinali, D.D. Venere and V. Linsalata, 2000. Role of endogenous flavonoids in resistance mechanism of Vigna to aphids. J. Agric. Food Chem., 48: 5316-5320.
CrossRefDirect Link - Li, B.Q., T. Fu, D. Yao, J.A. Mikovits, F.W. Ruscetti and J.M. Wang, 2000. Flavonoidbaicalin inhibits HIV-1 infection at the level of viral entry. Biochem. Biophys. Res. Commun., 276: 534-538.
CrossRefDirect Link - Culbreath, A.K., J.W. Todd and S.L. Brown, 2003. Epidemiology and management of tomato spotted wilt in peanut. Ann. Rev. Phytopathol., 41: 53-75.
CrossRefDirect Link - Holbrook, C.C., P. Timper and A.K. Culbreath, 2003. Resistance to Tomato spotted wilt virus and root-knot nematode in peanut interspecific breeding lines. Crop Sci., 43: 1109-1113.
Direct Link - Hurt, C.A., R.L. Brandenburg, D.L. Jordan, G.G. Kennedy and J.E. Bailey, 2005. Management of spotted wilt vectored by Frankliniella fusca (Thysanoptera: Thripidae) in Virginia market-type peanut. J. Econ. Entomol., 98: 1435-1440.
CrossRefDirect Link - Sobolev, V.S., B. Guo, C.C. Holbrook and R.E. Lynch, 2007. Interrelationship of phytoalexin production and disease resistance in selected peanut genotypes. J. Agric. Food Chem., 55: 2195-2200.
CrossRef - Jassim, S.A.A. and M.A. Naji, 2003. Novel antiviral agents: A medicinal plant perspective. J. Applied Microbiol., 95: 412-427.
CrossRefPubMedDirect Link - Wang, K.H., Y.H. Lai, J.C. Chang, T.F. Ko, S.L. Shyu and R.Y.Y. Chiou, 2005. Germination of peanut kernels to enhance resveratrol biosynthesis and prepare sprouts as a functional vegetable. J. Agric. Food Chem., 53: 242-246.
CrossRefDirect Link - Wang, M.L., A.G. Gillaspie, J.B. Morris, R.N. Pittman, J. Davis and G.A. Pederson, 2008. Flavonoid content in different legume germplasm seeds quantified by HPLC. Plant Genet. Resour., 6: 62-69.
CrossRef








