Research Article
Studies on Acid Stress Tolerant Proteins of Cyanobacterium
Department of Agricultural Microbiology, AC and RI, Madurai-625104, India
G. Gopalaswamy
Department of Agricultural Microbiology, AC and RI, Madurai-625104, India
INTRODUCTION
Cyanobacteria are photosynthetic prokaryotes that utilize oxygen evolving photosynthesis similar to higher plants (Brynt, 1987). Many cyanobacteria fix nitrogen under aerobic conditions in specialized cells called heterocysts which comprise 5-10% of cells in a filament (Fleming and Haselkorn, 1973). The unique feature of cyanobacteria is their versatility of occurrence (Brock, 1973). Cyanobacteria have been isolated from diverse habitats including saline soils (Amsaveni, 1995), hot springs (Ward et al., 1997) and coastal swamps (Komarek, 1998). The practice of utilizing cyanobacteria as an efficient source of biofertilizer for rice has been advocated and adopted in India (Venkataraman, 1981). Among the various soil factors affecting the distribution of cyanobacteria, pH is particularly important directly influencing cyanobacterial distribution and abundance in soil (Sardeshpande and Goyal, 1981). Cyanobacteria generally prefer neutral to slightly alkaline conditions (Singh, 1978).
Do acid-tolerant cyanobacteria exist? In a phytoplankton survey of 10 lakes in the Bavarian Forest as well as the lignite mining districts of Bavaria (Upper Palatine) and Lusatia, covering a pH gradient from 8.0 to 2.8, Steinberg et al. (1998) demonstrated that acid-tolerant cyanobacteria do exist. Later, Madhosoodhan and Dominic (1999) found that a large diversity of cyanobacteria are distributed profusely over a wide range of acidic pH regions of Kerala soils. Recently several promising cyanobacterial cultures were isolated from problem-specific soils in Tamil, Nadu India, which could definitely increase rice production in those soils. In India acid soils constitute a major component deficient in available nutrients. Only iron and aluminum are available in excess amounts but this causes toxicity to many microorganisms (Das, 1996). Previously Moore (1963) reported that many cyanophyceae were strongly related to soil acidity although acidity was a limiting factor. He also reported many nitrogen-fixing cyanobacteria in fairly acid soils. Aiyer (1965) reported that in acid soils many cyanobacteria flourish well even at an acidity of pH 3.8. However the predominant genera were recorded only in neutral or slightly alkaline soils (Fogg et al., 1973; Brock, 1973; Singh, 1978; Raghava Reddy et al., 1980). Jurgensen and Davey (1968) reported that cyanobacteria also survive under acidic conditions in field soils suggesting that they tolerate low pH levels. Gopalaswamy et al. (2002a) studied the biomass and biochemical constituents of acid tolerant cyanobacterium and found maximum acid tolerance at pH 5.
Stress alleviation in cyanobacteria has been known to be achieved through the production of stress proteins (Webb and Sherman, 1994). Cyanobacteria are known to synthesize a variety of proteins in response to various stresses. The mechanism of tolerance to various stresses is governed by the genes and the proteins assisting them. The exact mechanism of acid tolerance in cyanobacteria is yet to be elucidated. In the present investigation attempts were made to elucidate these specific proteins governing for acid tolerance mechanism from isolated acid tolerant strains of cyanobacterium. The proteins thus conferring the acid stress tolerant property were identified, purified and characterized. The acid tolerant protein so identified will be used to identify the specific gene that will pave the way for future transformation using other promising and predominant neutrophilic and alkalophilic cyanobacteria thereby making them a good candidate biofertilizer for all type of rice soils.
MATERIALS AND METHODS
Bacterial Strains
The present investigation was conducted in Algal Biotechnology Lab, Department of Plant Molecular Biology and Biotechnology, Tamil Nadu, Agricultural University (TNAU), Coimbatore, India during the period of June 2003 to October 2005. The cultures were isolated earlier, characterized and maintained in the laboratory. Six cyanobacterial cultures viz., Westiellopsis-ARM 48 (non acid tolerant culture), Westiellopsis AT-TGK-4A2 (acid tolerant), Westiellopsis-AT-TGK-4A7 (acid tolerant), Westiellopsis-AT-TGK-5B2 (acid tolerant), Nostoc-GG-SK-A7 (acid tolerant) and Anabaena-GG-SK-A7 (reference) were taken for the study.
Culture Conditions
The pure cyanobacterial cultures were inoculated at the rate of 0.00125-0.00150 g mL-1 sterile N-free BG-110 broth kept in 250 mL conical flasks. Seven days old cyanobacterial cultures were inoculated in all the flasks. The flasks were incubated under 3000 lux light intensity at 28±1°C, for 16/8 h day/night cycle for a period of 21 days. The cultures were grown on different pH of 4.0, 5.0, 6.0 and neutral condition.
Stress Treatments
The cultures in the flask were maintained to be in acidic medium using citrate buffer. The addition of the buffer were done regularly at an interval of 16 h, since the secretion of ammonia in the medium by the culture turns over the condition from acidic towards the normal/saline condition. The amount thus added for different pH was also standardized during the period of experimentation.
Sample Preparation and Protein Analysis
Total proteins were extracted from the acid tolerant and reference cultures. The total protein content of the cyanobacterial cultures was estimated by following the method of Lowry et al. (1951). The separation of protein was done by SDS-PAGE following the method described by Maniatis et al. (1987). Fifteen microgram of total protein were loaded onto each well to study the protein profile of the cultures.
Elution and Concentration
The size of molecular weight of the unknown band was determined using graphical method employing calibration curve (Weber and Osborn, 1969). To excise the identified protein of 15.7 kDa size and elute the protein from gel the electro elution method was adopted as described by Maniatis et al. (1987). The eluted samples in the vials were lyophilized based on Halliday and Baker (1985).
Blotting and Sequencing
The concentrate protein sample was ran in SDS-PAGE and were kept without staining and destaining for transferring to the PVDF membrane. The protein was blotted onto PVDF membrane (Pal corporation) using the Semi-dry blotting apparatus (Bio-Rad, USA). The N-terminal amino acid sequence was determined using the facility available at the Rajiv Gandhi Centre for Biotechnology, Thiruvananthapuram, Kerala following the Edman degradation method (Niall, 1973).
The analysis was carried out on a Shimadzu model PPSQ-21 protein sequencer using a Wako pack -PTH amino acid column (C18) maintained at 40°C. The coupling and conversion reactions were carried out at 45 and 65°C, respectively.
RESULTS
Cyanobacterial Protein Estimation
The total protein content of the standard reference cultures (adapted to neutral pH) and the acid tolerant cyanobacterial cultures was estimated 21 days after inoculation by Lowry et al. (1951) (Table 1).
The protein content of both acid and non acid tolerant cyanobacterial cultures varied significantly based on their tolerance to their pH. A higher protein content of 1280 µg mL-1 was observed in non acid tolerant Westiellopsis ARM 48 which drastically reduced to pH with 1090 µg mL-1. This shows that the growth and tolerance of Westiellopsis ARM 48 is affected by shift in pH towards acidity. A reverse trend was observed in acid tolerant cyanobacterial cultures which showed increased total content of protein towards the shift of acidity except Anabaena-GG-SK-A7. Since this culture can grow well at pH of 6.0 and gets reduced towards acidity. The protein content was 1720 µg mL-1 at pH 6.0 and reduced to 920 µg mL-1 at pH 4.0. At pH of 4.0 the highest total protein content was observed in acid tolerant Westiellopsis-AT-TGK-4A2 followed by Westiellopsis-AT-TGK-4A7, Nostoc-GG-SK-4B3 and Westiellopsis-AT-TGK-5B2, respectively. At pH of 7.0, the highest protein content was observed in Anabaena-GG-SK-A7 (Reference) and non acid tolerant Westiellopsis ARM 48 cultures.
From the above results it`s clear that acid tolerant cultures showed a higher total protein content at acidic pH and reduced towards salinity whereas non acid tolerant cultures showed increased protein content in neutral while comparatively reduced towards the shift of acidity.
Comparison of Non Acid Tolerant and Acid Tolerant Proteins of Cyanobacteria
About 15 µg of protein extracts from the acid tolerant and non acid tolerant cyanobacterial cultures derived from different pH were loaded in each well and the resulted profiles are presented in Fig. 1-6.
The comparison studies revealed no particular difference among the profile except that for non stress proteins, wherein unique band of 23 kDa was present which was absent in acid tolerant Westiellopsis- AT-TGK-4A2. These results revealed that these proteins govern specially for non acid tolerant condition. The cyanobacterial culture Anabaena-GG-SK-A7 was used as a reference culture for the study. In this present study, similar protein of 52±2 kDa was found to occur in Anabaena-GG-SK-A7 under acidic condition. Under similar conditions, Nostoc-GG-SK-A7 expressed a protein of same size. This protein was not expressed in other cultures viz., Westiellopsis-ARM 48, Westiellopsis-AT-TGK-4A2, Westiellopsis-AT-TGK-4A7 and Westiellopsis-AT-TGK-5B2. The comparison studies revealed no particular difference among the profile except that for non stress proteins, wherein one unique band of 23 kDa was present, which was absent in Anabaena-GG-SK-A7. These results were found to be similar for these proteins as in Westiellopsis-AT-TGK-4A2. These results confirm that the proteins govern specially for non acid tolerant condition. On subjecting Westiellopsis- AT-TGK-4A7 to acidic stress condition, induction of 15.7 kDa acid tolerant protein was found to be expressed. The same size proteins were also found to occur in Nostoc-GG-SK-A7 and Westiellopsis-AT-TGK-5B2. This protein was absent in other cultures. The result revealed the presence of a unique 15.7 kDa acid tolerant protein, which was absent in non-acid Westiellopsis-ARM 48. The result also revealed the presence of two more unique bands of size ranging 22-23 kDa that was absent in non-acid Westiellopsis-ARM 48. The results revealed the presence of a unique acid tolerant protein of size ranging 52±2 kDa in size also found in Anabaena-GG-SK-A7, which were absent in the non-acid Westiellopsis - ARM 48. The profile also revealed the presence of unique acid tolerant proteins of size ranging from 102-103 kDa. The presence of all this unique proteins confers that these proteins may govern for acid tolerance in acid tolerant cyanobacteria. On subjecting Westiellopsis-AT-TGK-5B2 to acidic stress condition, induction of 15.7 kDa acid tolerant protein was found to be expressed. The result also revealed the presence of unique band of size ranging 29±2 kDa. These proteins were found to be absent in non acid tolerant Westiellopsis-ARM 48.
| Table 1: | Total protein content of the non-acid stress and acid stress cyanobacteria (µg mL-1) |
 | |
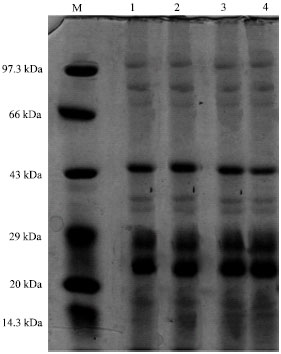 |
| Fig. 1: | Protein profile of Westiellopsis-ARM-48 at different pH, M: Medium range marker, Lane 1: Westiellopsis-ARM 48 at pH 4.0, Lane 2: Westiellopsis-ARM 48 at pH 5.0, Lane 3: Westiellopsis-ARM 48 at pH 6.0, Lane 4: Westiellopsis-ARM 48 at pH 7.0 |
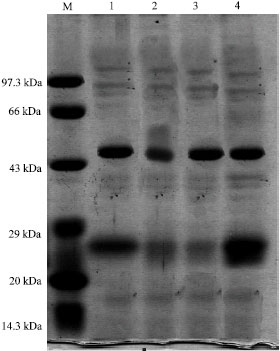 |
| Fig. 2: | Protein profile of Westiellopsis AT-TGK-4A2 at different pH, M: Medium range marker, Lane 1: Westiellopsis-AT-TGK-4A2 at pH 4.0, Lane 2: Westiellopsis-AT-TGK4A2 at pH 5.0, Lane 3: Westiellopsis-AT-TGK-4A2 at pH 6.0, lane 4-Westiellopsis-AT-TGK-4A2 at pH 7.0 |
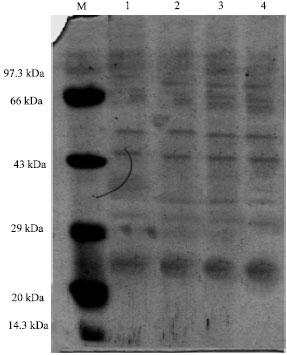 |
| Fig. 3: | Protein profile of Anabaena-GG-SK-A7 at different pH, M: Medium range marker, Lane 1: Anabaena GG-SK-A7 at pH 4.0, Lane 2: Anabaena GG-SK-A7 at pH 5.0, Lane 3: Anabaena GG-SK-A7 at pH 6.0, Lane 4: Anabaena GG-SK-A7 at pH 7.0 |
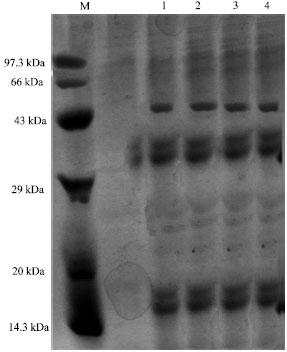 |
| Fig. 4: | Protein profile of Westiellopsis At-TGK 4A7 at different pH, M-Medium range marker, Lane 1: Westiellopsis At-TGK-4A7 at pH 4.0, Lane 2: Westiellopsis At-TGK-4A7 at pH 5.0, Lane 3: Westiellopsis At-TGK-4A7 at pH 6.0, Lane 4: Westiellopsis At-TGK-4A7 at pH 7.0 |
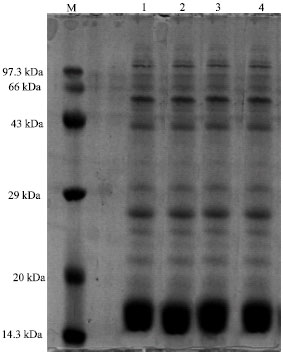 |
| Fig. 5: | Protein profile of Nostoc-GG-SK-4B3 at different pH, M: Medium range marker, Lane 1: Nostoc-GG-SK-4B3 at pH 4.0, Lane 2: Nostoc-GG-SK-4B3 at pH 5.0, Lane 3: Nostoc GG-SK-4B3 at pH 6.0, Lane 4: Nostoc-GG-SK-4B3 at pH 7.0 |
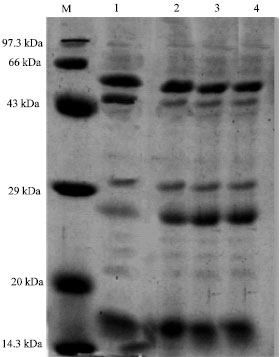 |
| Fig. 6: | Protein profile of Westiellopsis At-TGK 5B2 at different pH, M: Medium range marker, Lane 1: Westiellopsis-At-TGK-5B2 at pH 4.0, Lane 2: Westiellopsis-At-TGK5B3 at pH 5.0, Lane 3: Westiellopsis-At-TGK-5B3 at pH 6.0, Lane 4: Westiellopsis-AT-TGK-5B3 at pH 7.0 |
| Table 2: | N-terminal amino acid sequence of the acid tolerant protein |
 | |
| Single letter notation T F T S P P P P P Q. Three letter notation, Thr, Phe, Thr, Ser, Pro, Pro, Pro, Pro, Pro, Gln | |
| Table 3: | Degenerate nucleotide sequence for acid tolerant protein |
 | |
N-Terminal Amino Acid Sequence
The N-terminal amino acid sequencing of the 15.7 kDa protein was done for ten cycles and the resultant amino acid sequence is shown in Table 2. On back translation of the ten amino acids, a degenerate nucleotide sequence was obtained and is shown in Table 3.
DISCUSSION
Does an acid tolerant cyanobacterium exist? This remained a major contradiction and challenging perspective in phycology. As a result the occurrence of the phytoplankton mass was reported in acidic lake at pH of 8.0 to 2.8 (Steinberg et al., 1998). In a study conducted among the 240 composite samples 68, 89, 65 and 18 respectively, were reported from pH range 6-5, 5-4, 4-3 and 3-2.8 (Madhosoodhan and Dominic, 1999). Later few reports have been made on the isolation of cyanobacteria from acidic soils of India ( Gopalaswamy et al., 2002a). Few off cyanobacterium cultures were studied for their growth and biomass production and were found to be best at pH 4.0 (Gopalaswamy et al., 2002b). This confirms that cyanobacteria do live on acidic condition but the exact molecular mechanism remains yet to be revealed. Stress alleviation in cyanobacteria has been known to be achieved through the production of stress proteins (Webb and Sherman, 1994).
Huang et al. (2002) reported that Lethal acid stress at a pH below 4.4 results in the formation of aggregates of denatured proteins observed as granules near the cell periphery the disruption of the transmembrane pH gradient cell color change to blue and damage to photosystem II. In support to this in non acid tolerant Westiellopsis-ARM 48 only fewer protein were found to occur. There was much variation in other acid tolerant cyanobacterial cultures which produced other protein(s). But a unique band of 23 kDa was present which was absent in Westiellopsis-AT-TGK-4A2 that showed chlorosis at pH 7.0. This might infer that the specific protein may govern for saline tolerance mechanism. From the above result much of the present study can be correlated with various protein observed during stress conditions. On subjecting Westiellopsis-AT-TGK-4A7 to acidic stress condition induction of 15.7 kDa acid tolerant protein was found to be expressed. The same size proteins were also found to occur in Nostoc-GG-SK-A7 and Westiellopsis-AT-TGK-5B2. Roy et al. (1999) observed a similar trend in 16 kDa protein upon heat-shock treatment in the thermophilic cyanobacterium Synechococcus vulcanus.
Also a chloroplastic drought-induced stress protein 32 kDa was observed when induced by environmental and oxidative stress (Broin et al., 2002). The protein can be accorded to the protein of 29±2 kDa found in Westiellopsis-AT-TGK-5B2 specifically due to acid stress condition. The present study also revealed the presence of 52±2 kDa in Anabaena-GG-SK-A7 acidic condition under acidic condition. Under similar conditions Nostoc-GG-SK-A7 expressed a protein of same size. Similarly stress responsive protein of about 58 kDa was reported by Weissman et al. (1996) in Synechococcus sp. strain PCC 7942 in response to light temperature and metal stress. Eriksson et al. (2001) characterized clpB gene that encodes a 97 kDa protein with novel features in Synechococcus sp. strain PCC 7942 under heat shock. Recent study by Akira et al. (2001) revealed protein SynNhaP with a molecular weight of 129 kDa was in Synechocystis sp. PCC 6803 when subjected to an acidic pH of 5.0 in Synechocystis sp. PCC 6803. A similar trend was observed in the present study wherein a 103±2 kDa protein was found to occur at the same range in acid tolerant Nostoc-GG-SK-A7. Thus it is clear that this protein might be in response to the acidic stress condition. The above results confirm that the above revealed protein(s) are produced due to stress and govern for acid tolerance mechanism.
The mechanism of tolerance to various stresses is governed by the genes and the proteins assisting them. With the help of derived a degenerate nucleotide, primer sequence can be constructed to identify the desired fragment or gene governing for acid tolerance. This may be designed that could pave the way for the cloning and sequencing of the gene encoding the acid stress protein. The gene could be cloned and used for transformation to other promising and predominant neutrophilic and alkalophilic cyanobacteria thereby making them good candidate biofertilizers for all types of rice soils for wider adaptability.
ACKNOWLEDGMENTS
We acknowledge our sincere thanks to Rajive Gandhi centre for biotechnology for providing us the N-terminal sequencing. We also thanks School of Post Graduate Studies (SPGS), TNAU for providing fund to successfully complete the part of research work.