Research Article
Sequence Analysis of a Comparative Anchored Tagged Sequence (Biglycan) and its Assignment to the X Chromosome of River Buffalo
Department of Cell Biology, Genetic Engineering Division, National Research Center, Dokki, Giza, Egypt
Genomes of domestic animals are practically unknown when compared with those of both humans and mice. Cattle is considered to be the most studied bovid species, where 4,357 loci are mapped with 1,507 being genes (INRA bovmap database, July, 2006). They are mostly mapped to large chromosome regions. Another important bovid species is the river buffalo (Bubalus bubalis, 2n = 50, BBU), where more than 130 million of river buffaloes are raised all in the world for both meat and milk production. With its economic importance in many Asian and Mediterranean countries, the genetic improvement of river buffalo, especially in reproductive performance and quantity of meat and milk production, ranks high among agricultural research needs of these countries. Genome studies and a physical gene map of river buffalo is being developed and expanding.
Comparative mapping, mapping homologous genes in multiple species, provides more information about chromosomal evolution between distant mammalian species than is available from the best cytogenetic technologies currently available. In this respect, the rapid development of the human and mouse gene maps has stimulated the expansion of genetic maps for several additional mammalian species. A common observation, which became the basis for the field of comparative genome mapping, is the remarkable conservation of linkage organization of homologous genes in species from diverse mammalian orders (Copeland and Jenkins, 1991; Womack and Kata, 1995; Johansson et al., 1995; Nadeau, 1995).
Precise comparisons of mammalian gene maps require common anchor loci as landmarks for conserved chromosomal segments. In this respect, Lyons et al. (1997) designed a number of evolutionarily conserved primer pair sequences termed Comparative Anchor Tag Sequences (CATS), providing PCR-format gene markers that can be used to construct and connect gene maps of any mammalian species.
Studies on the use of CATS primers in river buffalo were initiated at the Department of Cell Biology, National Research Center, Cairo, Egypt and Department of Veterinary Pathobiology, College of Veterinary Medicine, Texas A and M University, USA, through a joint project (Bio-005-002). Biglycan (BGN) is defined to be an extracellular matrix molecule that is important for the maintenance of muscle cell integrity (Mercado et al., 2006). In the present study, primer pair representing BGN and published as CATS by Lyons et al. (1997) was used to investigate its reactivity in river buffalo and to study its segregation in a panel of 43 somatic cell hybrids, obtained from the fusion of river buffalo blood lymphocytes and cells from Chinese hamster cell line (wg3h) (de Hondt et al., 1991), in an attempt for the assignment of BGN to river buffalo chromosomes. The obtained results necessitate sequencing of the amplified PCR product of buffalo DNA and Blast search of Genbank to verify the BGN assignment.
The PCR primer pair used in this study for BGN was one of the CATS designed by Lyons et al. (1997). The sequences of the forward and reverse primers are: CTC CAA GAA CCA CCT GGT GTTC AAA GCC ACT GTT CTC CAG A panel of 43 somatic cell hybrids resulting from the fusion of river buffalo lymphocytes and Chinese hamster cell line wg3h (de Hondt et al., 1991), was used to investigate the segregation pattern of BGN anchored locus by Polymerase Chain Reaction (PCR).
Genomic DNA extraction: The isolation of DNA from buffalo leukocytes was performed according to Blin and Stafford (1976), while hamster cell line and somatic cell hybrids DNA was extracted according to Ausubel et al. (1990).
Buffalo blood was collected in syringes containing Ethylene Diamine Tetra Acetic Acid (EDTA). Ten mL blood were transferred to a 50 mL polypropylene tube on ice to which 25 mL of cold 2X sucrose-Triton (0.64M sucrose; 0.02 M Tris-base; 0.01M MgCl2; 2% Triton X-100 pH 7.6) and 15 mL dd H2O were added. The tube was inverted a few times and set on ice for 10 min with occasional inversion. The mixture was centrifuged at 5000 rpm for 15 min at 4°C and the supernatant was discarded. The cultured cells of hamster and hybrid cell lines were scrapped, centrifuged at 2000 rpm for 15 min at room temperature and the supernatants were discarded.
The cell pellets from buffalo blood, hamster and hybrid cells were suspended in 3 m nuclei lysis buffer, in addition to 1/20 volume of 20% SDS and 1/20 volume of proteinase K(10 mg mL-1). The samples were incubated overnight in a shaking water-bath at 37°C.
DNA from each sample was extracted once with TE-saturated phenol, then with phenol: chloroform: isoamyl alcohol (25: 24: 1) until there was no protein at the interface and finally by chloroform-isoamyl alcohol (24:1). For each extraction, the aqueous phase was mixed well with an equal volume of solvent, centrifuged for 10 min at 2000 rpm and the top layer carefully transferred to another falcon tube for the next extraction. To the final aqueous phase, 1/10 the volume of 2.5 M NaOAc (pH 5.5) and 2.5 volume of cold 95% ethanol were added, the tube was agitated to mix. DNA was picked up with a heat-sealed Pasteur pipette and washed briefly in cold 70% ethanol and air dried, then dissolved in an appropriate volume of 1X TE buffer. DNA concentration was determined using UV spectrophotometer (Pharmacia LKB-Ultrospec) at 260 nm. A reading of 1.0 at 260 nm is equivalent to 50 μg mL-1.
Polymerase chain reaction: PCR was performed in 25-100 μL reaction mixture consisting of 0.2 mM dNTPs, 10 mM Tris, 50 mM KCl, 1.0 to 2.0 mM MgCl2, 1% Triton X-100, 2.5 units Taq polymerase and 1.0 μM forward and reverse primers, this was aliquoted into PCR tubes with 100 mg DNA of buffalo, hamster or hybrid cells. The reaction mixture was overlaid with sterile mineral oil and was cycled in the Techne Cyclogene PCR machine using 1 cycle (3 min) at 94°C, followed by 30 cycles (1 min at 94°C, 1 min at 64°C and 1 min at 72°C) and finally 1 cycle (10 min) at 72°C. Following the completion of the PCR cycles, 2 μL of tracking dye (20 mg bromophenol blue, 1 mL 0.2 M EDTA, 2 mL glycerol, sterile distilled water to 20 mL total volume) was added to each PCR tube. The reaction products and appropriate size marker were electrophoresed on 1.5% agarose in 1X-Tris acetate buffer (TAE) containing 0.8% μL of 10 mg mL-1 ethidium bromide. After electrophoresis, the gel was examined with UV at a wave length of 312 nm and photographed using a Polaroid MP4+ camera.
Statistical analysis: The segregation pattern of the PCR product was compared with bovine synteny groups and X chromosome. Synteny was determined by calculating the correlation coefficient (φ) according to Chevalet and Corpet (1986). A φ value of more than 0.67 is indicative of synteny.
Sequence analysis: The PCR product of river buffalo DNA was purified and sequenced at the Center of Genetic Engineering, Ain Shams University/Cairo/Egypt. Sequence analysis and alignment were carried out using NCBI-BLASTN 2.2.14 version (Altschul et al., 1997).
In order to assign BGN marker, as one of the CATS, to river buffalo; the primer pair representing the marker under investigation was tested for reaction with both buffalo and hamster cell line DNAs, where it was found to react relatively with buffalo only and after optimization of the annealing temperature, an intensive PCR product of approximately 700 bp was obtained.
A panel of 43 buffalo/hamster somatic cell hybrids was used for gene assignment, where they were tested for reactivity with BGN primer pair.
The presence or absence of PCR amplified product was investigated in the hybrid panel. The segregation pattern of the studied marker was compared with that of biochemical or molecular markers representing all bovine synteny groups analyzed using the same hybrid panel and already assigned to buffalo chromosomes.
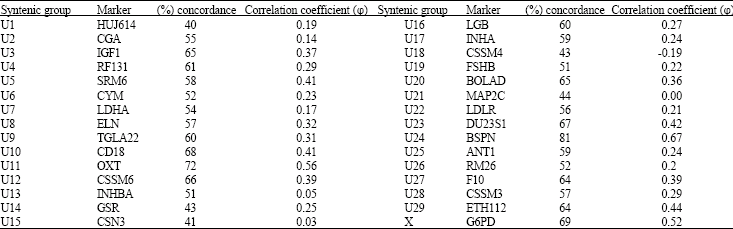
Synteny was determined by calculating the percent concordance and the correlation coefficient (φ) (Table 1). Statistical analysis showed syntenic relation with U24 (φ = 0.67). It also gave a rather high φ value with U11 (φ = 0.56) and with markers representing the X chromosome (φ = 0.52).U24 has been assigned to cattle chromosome 14 (BTA14) as reported by Barendse et al. (1994), which is homologous to buffalo chromosome 15 (BBU15) (CSKBB, 1994). Therefore the results suggest that BGN can be assigned to BBU15.
Since BGN has been assigned to the X chromosome in cattle (Wegrowski et al., 1995; Amaral and Womack, 2000; Rubes et al., 2005) and human (Mc Bride et al., 1990; Traupe et al., 1992) and also because of the possible presence of unexpected non-homologous amplification in different species, the results required further investigations. A one way sequencing of the BGN amplified PCR product of buffalo DNA (amplicon) was performed using the reverse primer, where the buffalo amplicon obtained was found to be 636 bp (Fig. 1). Sequence analysis and alignment were carried out using NCBI-BLASTN 2.2.14 version (Altschul et al., 1997). The Blast searches of GenBank revealed that the amplified PCR segment shares sequence with Bos taurus chromosome X genomic BGN with 95% identities (GenBank accession numbers NW001030256.1/BTX WGA42622). The shared sequence is extending from the buffalo amplicon’s nucleotide number 44 to 635 and the BTA (BGN) nucleotides 298302 to 298893. Eight gaps were observed in this alignment, specifying amplicon’s nucleotides number 67, 71, 86 and 451 and BTA (BGN) nucleotides number 298363, 64, 65 and 298885 (Fig. 2A). The buffalo amplicon was also found to share sequence with two segments of Homo sapiens chromosome X (HSA X) genomic BGN with 91% identities each (gi|22773406|gb|U82695.3| and gi|22773273|gb|AF274858.3|). The shared sequence in segment 1 (Fig. 2B) extended from the buffalo amplicon’s nucleotide number 497 to 626 and from HSA (BGN) nucleotides 110819 to 110690. Whereas in the second segment, the shared sequence was found to extend from nucleotide number 497 to 626 of the amplicon and from HSA (BGN) nucleotides 65467 to 65338, with no gaps observed in neither segments 1 nor 2 (Fig. 2C).
 | |
| Fig. 1: | Nucleotide sequence of BGN amplicon |
| Table 1: | Percent concordance and correlation coefficient (φ) of BGN marker with markers for bovine syntenic groups and the x chromosome |
 | |
 | |
| Fig. 2a: | Sequence alignments between BGN amplicon and Bos Taurus (BTA) chromosome X genomic BGN |
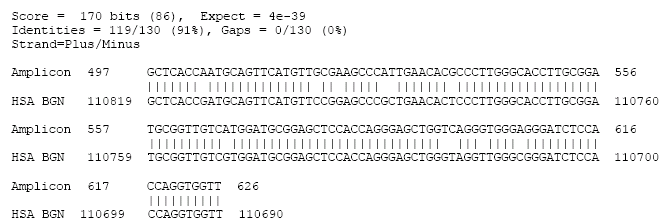 | |
| Fig. 2b: | Sequence alignments between BGN amplicon and Homo sapiens (HSA) chromosome X genomic BGN (segment 1) |
 | |
| Fig. 2c: | Sequence alignments between BGN amplicon and Homo sapiens (HSA) chromosome X genomic BGN (segment 2) |
Effective comparative mapping inference utilizing developing gene maps of animal species requires the inclusion of anchored reference loci that are homologous to genes mapped in the more gene-dense mouse and human maps. Nominated anchor loci, termed comparative anchor tagged sequences (CATS), have been designed by Lyons et al. (1997) to facilitate direct comparisons between divergent species gene maps, where they represent landmarks for conserved chromosomal segments and providing PCR-format gene markers that can be used to construct and connect gene maps of virtually any mammalian species.
Preliminary genetic maps for several mammalian species have been developed, primarily utilizing high resolution polymorphic microsatellite markers (Type II markers); however, these have little comparative value because they are seldom conserved outside the mammalian order of the studied species (Lyons et al., 1997). The best markers for such anchor loci would be expressed genes (termed Type I markers) as their DNA sequence can be used to establish homology between divergent species (O’Brien, 1991).
The assignment of molecular markers, especially type I markers, to domestic animal chromosomes represents an important goal for their genetic improvement. Relatively few loci have been mapped in river buffalo (Bubalus bubalis, 2n = 50) in comparison with those mapped in cattle (BovBase). However, considering the high degree of chromosome band homologies and loci conservation existing between cattle and river buffalo (CSKBB, 1994; De Hondt et al., 1997; De Hondt and El Nahas, 2001; Di Meo et al., 2000; El Nahas et al., 1993, 1996, 1997, 1998, 1999, 2001; El Nahta et al., 1994; Iannuzzi et al., 2000a, b; Mahfouz, 2000; Oraby et al., 1998; Othman and El Nahas, 1999; Othman et al., 2003; Othman and Bibars, 2004), data on the bovine physical map should help to extend those of related species, in particular river buffalo.
In the context of comparative mapping studies, a number of CATS consensus primers, from loci mapped to human chromosomes 9, 10, 20 and 22, have been used to amplify homologous loci in pigs. Ten loci were physically mapped where map locations were consistent with human/pig Zoo-FISH (Lee et al., 2001). Whereas, Hassan (2004) tested the reaction of one of the CATS (LDHB) on buffalo DNA and used a panel of Buffalo/hamster hybrid cells for gene assignment, where synteny of LDHB with two syntenic groups, U3 and U29, was reported earlier (El Nahas et al., 1999) which were assigned to the long and short arms of BBU4, respectively. The sequence analysis, conducted by Hassan (2004), of the buffalo amplicon revealed the assignment of an LDHB pseudogene to the X chromosome of river buffalo.
In this respect, some genes can be used to expand the physical map of river buffalo, by verifying the assignment of genes to chromosomes and by assigning new ones. This is the case in the present study, where BGN has not yet been assigned to river buffalo chromosomes. The primer pair for BGN gene reacted positively with river buffalo DNA producing a buffalo PCR product of approximately 700 bp. The same band size was also shown by Lyons et al. (1997), using the same primer pair and other mammalian DNAs.
The results in this study indicate syntenic relation of the investigated BGN marker and markers representing U24 (φ = 0.67). Synteny with U11 and the X chromosome had φ values of 0.56 and 0.52, respectively. It is to be noted that some unexpected chromosomal loss or rearrangements may occur in somatic cell hybrids resulting into misleading or confusing informations ; this drawback was overcome by sequence analysis of the amplified PCR product of buffalo DNA and Blast searches of GenBank, since the presence of unexpected non-homologous amplification in different species is possible. Sequence analysis revealed sequence homology between the buffalo amplicon and sequence of Bos taurus chromosome X genomic BGN with 95% identities (NW001030256.1/BTX WGA42622) and with two segments of Homo sapiens chromosome X genomic BGN with 91% identities each (gi|22773406|gb|U82695.3|and gi|22773273|gb|AF274858.3|).
Lyons et al. (1997) also reported the homology of the feline PCR product sequence of BGN with both sequences of Bos taurus and Homo sapiens.
BGN was previously reported to be assigned to Bos taurus chromosome X (BTA X) by Wegrowski et al. (1995), Amaral and Womack (2000) and Rubes et al. (2005). On the other hand, the localization of BGN gene to human chromosome X (HSA Xq13-qter) was reported by Mc Bride et al. (1990) and more precisely, fine mapping of BGN within the human Xq28 region was performed by Traupe et al. (1992) employing a hybrid cell panel. Whereas, genetic and physical mapping of BGN to the mouse X chromosome (MMU X) was accomplished by Chatterjee et al. (1993). BGN was also mapped to the X chromosome of the horse (Equus caballus), wallaby (Macropus eugenii), platypus (Ornithorhynchus anatinus), planigale (Planigale maculate sinualis) and rats (Rattus norvegicus) as referenced in the Horsemap database for LIFE for BGN. These reported assignments of BGN to different species chromosome X are in agreement with the fact that genes on the X chromosome have been conserved in many species, where human and mouse X chromosomes contain a minimum of eight conserved segments (Blair et al., 1994). The X chromosome is also conserved in human and cattle (Amaral et al., 2002). Markers of the X chromosome, previously reported to be assigned to river buffalo are conserved on human and cattle X chromosomes as is the case with G6PD (Mahfouz, 2000; Othman et al., 2003), PGK1 (Iannuzzi, 1998; Iannuzzi et al., 1999; Iannuzzi et al., 2000a) and F9 (Othman et al., 2003).
Despite the close relation and the strikingly similar banding patterns between buffalo and cattle, since they both belong to subfamily Bovinae (CSKBB, 1994), there is no doubt that some differences do exist between the two species; this is slightly observed in this study, where the alignment of the buffalo amplicon with Bos Taurus chromosome X BGN revealed 95% identities with 8 gaps observed. This difference in the nucleotide sequence between both species indicates the necessity of a new serge of investigations to be conducted in order to reveal the differences between buffalo and cattle since almost all the searches made so far were concentrating on the close relation and the similarity between both species.
In conclusion, comparative mapping predicts the conservation of the X chromosome in different mammalian species, but some internal chromosome rearrangements may exist among them, therefore the results presented here allow the assignment of BGN gene to BBU X for the first time, based on comparative mapping predictions and the subsequent investigations in this study, thus extending the river buffalo physical map and confirming the extensive genetic conservation between buffalo, cattle and human chromosomes especially for the X chromosome.
The author wishes to acknowledge professor S.M. El Nahas, Cell Biology Unit, National Research Center, for her sincere help and careful examination of the manuscript. This study was in part supported by the joint research project (Bio-005-002).