Research Article
Production, Partial Characterization and Cloning of Thermostable α-amylase of a Thermophile Geobacillus thermoleovorans YN
Alexandria, Egypt
Nadia A. Soliman
Alexandria, Egypt
Yasser R. Abdel-Fattah
Alexandria, Egypt
One of the most abundantly distributed polysaccharides in nature is starch, which is produced by plants. It is composed of two high molecular weight compounds, amylose and amylopectin. Amylose is a linear chain of glucose residues linked with an α-1,4 bond. Amylopectin is a branched polymer where the α-1,4-linked glucose residues are branched every 17-26 residues with an α-1,6-linked points. A wide variety of microorganisms are able to degrade and utilize this natural high molecular weight biopolymer by secreting starch-degrading enzymes. These enzymes work either from the non-reducing end of the chain acting as exo-enzymes producing low molecular weight products (i.e., β-amylase, glucoamylase and α-glucosidase) or in the interior of the chain and in a random fashion acting as endo-enzymes and producing linear and branched saccharides with various lengths (i.e., α-amylase). A great number of α-amylases (E.C. 3.2.1.1) have been isolated from a variety of eucaryotic and procaryotic organisms and described by Antranikian, (1990), Leuschner and Antranikian (1995). All of them have been compiled in family 13 of the classification of the glycosyl hydrolase superfamily described by Henrissat and Bairoch (1996). Two α-amylases from Dictyolglomus thermophilum (Fukusumi et al., 1988) and Pyrococcus furiosus (Laderman, 1993a, b), could not be classified in family 13 and have been included in newly established family 57 (Jorgensen et al., 1997).
Amylases constitute a class of industrial enzymes having approximately a 25% stake in the world enzyme market. Of these, α-amylase that plays a key role in starch conversion technology by making starch usable for other amylases. Among the starch-hydrolyzing enzymes that are produced on an industrial scale, thermostable α-amylases are of considerable commercial interest. Bacteria belonging to the genus Bacillus have been widely used for the commercial production of thermostable α-amylases due to their potential biotechnological uses in the food, pharmaceutical and fine chemical industries. These include α-amylase from Bacillus coagulans, B. stearothermophilus, B. caldolyticus, B. brevis, B. acidocaldarius and B. thermoamyloliquefaciens (Campbell, 1954, 1955). The most important characteristic of thermophilic organisms is their ability to produce thermostable enzymes with a higher operational stability and a longer shelf-life (Niehaus et al., 1999). As compared with eubacterial enzymes, archaebacterial amylases from Pyrococcus furiosus and P. woesei exhibit greater thermostability (Brown et al., 1990; Koch et al., 1990, 1991) which produce α-amylases active at a temperature of 100°C and above (Malhotra et al., 2000). Since these natural thermophilic isolates are not considered suitable for use in commercial processes because of their very low productivities and the high energy expenditure involved in growth and enzyme production (Sidhu et al., 1997), isolation of a strain is invariably followed by its improvement for production using mutagenesis and/or recombinant DNA technology and bioprocess optimization.
In this study we report the production of thermoactive and thermostable α-amylase from locally isolated lipase-producing Geobacillus thermoleovorans YN. Monitoring of enzyme production as well as SDS-PAGE and zymogram analysis of amylolytic activity was investigated. Emphasis was given to the biochemical characterization of the crude enzyme in terms of its optimal temperature and thermal stability. PCR cloning and sequencing of the gene responsible for α-amylase enzyme production was also described.
Bacterial strains and plasmids: The microorganism used in this research as a potent thermostable amylase producing strain, Geobacillus thermoleovorans YN (accession number AF385083), was isolated and characterized as described in (Abdel-Fattah et al., 2002). For cloning purpose, Escherichia coli DH5-α of the genotype [Sup E44 ΔlacU169 (Φ80lacZΔM15) hsdR17 recA1] (Hanahan, 1983) was used as host strain, where Bluescript ® II KS(+) (Stratagene, Inc.) was used as a cloning vector.
Growth and enzyme production: A preliminary test for production of α-amylase by Geobacillus thermoleovorans YN was carried out by cultivation of bacterial strain under shaking conditions (200 rpm) at 55°C on Luria-Bertani (LB) complex medium (M1) containing (w/v): Tryptone, 1%; yeast extract, 5 and NaCl, 5 or basal medium (M2) containing (w/v): Corn starch, 1%; yeast extract, 0.3%; (NH4)2SO4, 0.3%; K2HPO4, 0.1%; MgSO4. 7H2O, 0.02% and NaCl, 0.1% (pH adjusted to 7.0 before autoclaving). To study the effect of glucose on enzyme induction, bacterial strain was grown on basal salt medium without starch and supplemented with 1% glucose (M3) as the sole carbon and energy source. The production of enzyme was measured daily for 72 h.
Enzyme assays: Amylase activity was routinely estimated by measuring the reducing sugar released during the reaction, using starch as the substrate, according to Somogyi and Nelson (Nelson, 1944’). The reaction mixture contained 50 μL of 1.1% soluble starch (Riedel deHähn) in 2 mM imidazole-HCl buffer (pH 7.0) and 250 μL of enzyme solution. The reaction was stopped by adding 100 μL dinitrosalicylic acid solution (100 mL of solution containing 1 g 3,5-dinitrosalicylic acid, 30 g potassium sodium tartarate and 20 mL 2 N NaOH) after incubation at 60°C for 15 min incubation time. The reaction mixture was heated in boiling water for 5 min and the absorbance at 540 nm was measured after cooling in ice and diluting with 1 mL distilled water. A standard curve was prepared using different concentration of D-glucose and the absorbance was measured at 540 nm. One unit of enzyme was defined as the amount of enzyme that releases 1 μmol of reducing sugar from the substrate per minute under the condition of assay method.
Optimum temperature for enzyme activity was determined by measuring the activity in a temperature range 40-95°C. For testing the enzyme thermal stability, it was incubated at temperatures (75 and 80°C) for 5, 10, 15, 30, 45 and 60 min and the residual soluble starch digesting activity was assayed as described previously.
Gel electrophoresis and zymography: Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) was performed as described by Laemmli (1970). After electrophoresis, the gel was renatured by overnight incubation at 4°C in 50 mM phosphate buffer (pH 7.0) containing 2.5% Triton x-100. The gel was then washed twice in the same buffer without triton. The renatured gel was incubated in reaction solution containing 1% soluble starch, 20 mM sodium acetate (pH 5.5) and 1 mM CaCl2. The gel was rinsed with iodine solution (1% I2, 10% KI and 50% ethanol) (Kim et al., 1998). The amylolytic activity appears on the SDS gel as a colorless band.
DNA isolation, manipulation and PCR amplification: Preparation of genomic DNA from Geobacillus thermoleovorans YN was performed according to Ausubel et al. (1987). Preparation of plasmid DNA, digestion with restriction endonucleases, separation of fragments by agarose gel electrophoresis, ligation of DNA fragments were performed as described by Sambrook et al. (1989). Polymerase Chain Reaction (PCR) techniques were performed with Taq polymerase (Fermentas) and the amplified fragment was cut and purified from agarose gel using SephaglasTM BandPrep kit.
PCR-cloning: Polymerase chain reaction (PCR) technique was performed for amylase gene amplification using two sets of primers designed based on sequencing alignment of α-amylase producing thermophilic bacilli and geobacilli (accession numbers: AF438149, BD144004, BD144003, D90112, Y17557, J01547, AX453590, AX453588, AX453586, AX453584, AX370736, AX370734, AB051102, AF032864, E01181, E01180, E01157 and X02769). The first set amplified a 1538 bp consensus region, while the second set amplified 2146 bp from genomic DNA of the tested bacterial strain. The blunt end PCR fragment (2146 bp) was ligated to Bluescript ® II KS(+) cut with EcoRV.
Electroporation: Transformation of E. coli DH5-α competent cells was carried out by electroporation according to Sambrook et al. (1989). E. coli was grown in LB medium at 37°C. Cultures were diluted 1:10 into 25 mL of pre-warmed media and incubated with aeration until the cells reached early log phase (1x108 to 2x108 cells mL-1 equivalent to OD550 0.5-0.8). They were transferred to centrifuge tubes, incubated on ice for 15 min and kept cold through the rest of the procedures. The cells were pelted at 2,700xg for 10 min at 4°C and washed twice with 10 mL of 1 mM cold HEPES (pH 7.0). The cells were re-suspended in 5 mL of 10% cold glycerol. Aliquots of 65 μL were shock-frozen in liquid nitrogen and stored at -80°C. After thawing on ice, competent cells was mixed with blunt ligated mixture DNA (0.25 to 0.5 μg mL-1). The mixture was transferred to a cold 0.1-cm-diameter cuvette. One pulse with the Multiporator (Eppendorf, Germany) was set a 2.0 kV and 25 μF, with the pulse controller set at 200 Ω. The cells were immediately diluted with 0.9 mL of LB and incubated at 37°C for 1 h. The cells were plated onto transformants selective medium supplemented with 100 μg mL-1 ampicillin and incubated at 37°C. For experiments utilizing α-complementation, isopropyl-thio-β-galactoside (IPTG) and 5-bromo-4-chloro-3-doyl-β-D-galactoside (X-Gal) were added to LB agar media at concentration 100 and 20 mg L-1, respectively.
DNA sequencing and sequence analysis: The nucleotide sequence of the PCR products and cloned fragments was determined on both strands using the chain-termination method of Sanger et al. (1977), with specific and universal primers, respectively. Sequence analysis of the DNA fragments was done using BLAST family programs. The nucleotide sequence of the gene along with amino acid translation is illustrated in Fig. 4. Multi aligment between the target sequence and the closely related (ac: Y17557, X59476, AF032864, X02769, AY705090, M11450 and M57457) was performed with CLUSTAL W.
Molecular screening and selection of potent α-amylase thermophilic producing bacteria: In a program for molecular screening of α-amylase producing strain, PCR primers were designed according to alignment data of α-amylase nucleotide sequences of thermophilic bacilli and geobacilli as described previously. These primers were used to amplify a conserved region of the gene. The PCR reaction was carried out on genomic DNA of a number of previously isolated thermophilic bacilli, where only three isolates showed amplification fragment with molecular size of 1538 nucleotides. The most potent strain in respect to α-amylase production, as detected by starch agar plate method, was the previously isolated G. thermoleovorans YN and reported as thermostable lipase producing strain (Abdel-Fattah et al., 2002). On sequencing the PCR fragment, more than 99% similarity to α-amylases, specifically the maltohexaose-producing enzyme α-amylase, from Geobacillus stearothermophilus was measured (Ben Ali et al., 2001).
In order to test the capability of the potent lipase-producing Geobacillus thermoleovorans YN to produce α-amylase enzyme, a preliminary test was performed to test the level of enzyme production by this bacterial strain on different media. A 1% of 12 h old preculture cell in LB was used to inoculate the tested media (M1: LB complex medium, M2: medium containing starch as the sole carbon source and M3: medium containing glucose as the sole carbon source), then allowed to grow in shaking (200 rpm) at 55°C for 72 h. The α-amylase production by the investigated strain (Geobacillus thermoleovorans YN) was compared daily in these media. Cells were harvested and the supernatants were tested for α-amylase activity as described in Materials and Methods. Results in Table 1 showed that the level of enzyme activity in supernatant remains constant after 24 h, whereas a sharp drop in activity was shown after 72 h incubation. This is in accordance to Rothstein et al. (1986) who showed that α-amylase in B. licheniformis SA1 is produced predominantly during growth and not during the stationary phase.
| Table 1: | Production of α-amylase enzyme by Geobacillus thermoleovorans YN during growth on different media |
 | |
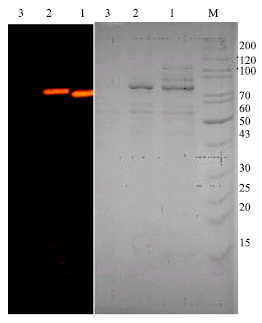 | |
| Fig. 1: | SDS-polyacrylamide gel electrophoresis and zymogram of produced α-amylase in different media after 24 h time incubation. The right gel was stained with Coomassie brilliant blue; the left gel illustrates in situ amylase activity detection. Lane M, standard protein as molecular weight (kDa); lane 1, amylase produced by G. thermoleovorans YN in M1; lane 2, amylase produced by G. thermoleovorans YN in M2; lane 3, amylase produced by G. thermoleovorans YN in M3 |
Approximately five times as much α-amylase was present when cells were grown in medium containing starch instead of glucose or LB complex medium (Table 1). These results do not reflect the inducible secretion of this enzyme but reflect the catabolic repression caused by glucose and the constitutive production of enzyme as described by Malhotra et al. (2000).
On the other hand, SDS-PAGE and zymogram of cell free supernatant from the three different types of media was performed (Materials and Methods). The cell free supernatants were developed from cultivation of G. thermoleovorans YN at 55°C in different media. Results shown in Fig. 1 indicated that the active amylolytic protein with apparent molecular weight around 62 kD.
Effect of temperature on Geobacillus thermoleovorans amylase activity: In order to test the effect of temperature on the α-amylase enzyme activity produced by Geobacillus thermoleovorans YN. The activity of enzyme was determined after incubation of the crude enzyme preparation, during assay conditions, with starch substrate under different temperatures ranging from 40 to 95°C. Results in Fig. 2 showed that the optimal enzyme activity was recorded at 75°C, while less than 50% of the optimal activity was measured under 60°C.
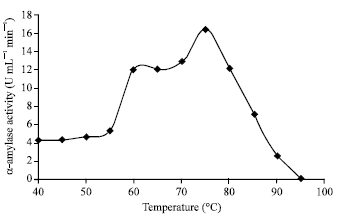 | |
| Fig. 2: | Effect of temperature on α-amylase enzyme activity produced by G. thermoleovorans YN |
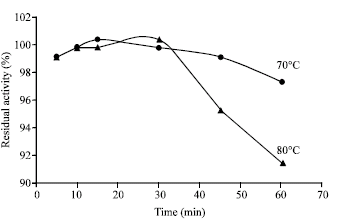 | |
| Fig. 3: | Thermal stability of α-amylase enzyme activity produced by G. thermoleovorans YN |
These results indicate the thermophilic nature of the enzyme. Increasing the reaction incubation temperature over 75°C led to exponential decrease in enzymatic activity. On the other hand the reported starch hydrolysis enzyme of Thermus sp. (Shaw et al., 1995), Bacillus sp. WN11 (Mamo et al., 1999), Bacillus sp. (Ben Ali et al., 1999), Bacillus licheniformis (Tsurikova et al., 2002) Bacillus thermoleovorans NP54 (Malhotra et al., 2000) worked optimally at 70, (75-80), 82, (90-95) and 100°C, respectively.
Thermal stability of α-amylase from Geobacillus thermoleovorans YN: One of the key factors determines the applicability of the α-amylase enzyme for industrial processes is its thermal stability. For this reason, the stability of α-amylase produced by Geobacillus thermoleovorans YN was tested by determination of the residual enzyme activity after heat treatment of the crude enzyme preparation by incubation at 75 or 80°C at a different time intervals ranging from 5 to 60 min. Results shown in Fig. 3 indicated that the enzyme optimally stable at 70°C over 60 min retaining more than 95% of its activity, while the highest loss of activity was obtained (1.05 fold decrease) after exposure of enzyme to 80°C for 60 min.
 | |
 | |
 | |
| Fig. 4: | Nucleotide sequence of the amylase from G. thermoleovorans YN. The predicted amino acid sequence is given below the nucleotide sequence in the standard one-letter code. The ORFs are bold and the stop codon is marked by an asterisk. The solid underlined lines expressed the positions of the first primer set, the dotted underlined lines expressed the positions of the second primer set and the arrows indicate the primer’s directions |
These results collectively indicated that the enzyme is a thermophilic and showed a complete thermal stability at 80°C for 30 min. Among bacilli and geobacilli capable of producing α-amylase, there is a great discrepancy in the thermostability of enzyme produced that related to strain. Bacillus sp. US100 showed a remarkable thermostability with a half life of 40 min at 110°C (Ben Ali et al., 1999) and Bacillus subtilis appeared very stable since more than 80% of this activity was still left after 2 h of incubation at 90°C (Konsoula and Liakopoulou-Kyriakides, 2007).
The recorded significant thermostability of α-amylase produced by G. thermoleovorans YN the strain used in this work make it a good candidate for industrial applications.
Molecular cloning of the gene encoding α-amylase activity of Geobacillus thermoleovorans YN in E. coli DH5-α: Further molecular characterization of the α-amylase enzyme was performed by PCR amplification of the whole gene. Taking the advantage of close similarity of the partial sequence of B. thermoleovorans YN α-amylase gene with the sequence of maltohexaose-producing enzyme α-amylase, from Geobacillus stearothermophilus, set of PCR primers were designed (2 pairs of primers were designed) and used in PCR technique. The longer amplified blunt end PCR fragment (2146 bp) was cloned by ligation to Bluescript ® II KS(+) vector cut with EcoRV, transformed into E. coli DH5-α competent cells by electroporation and screened on LB-agar plates induced with IPTG. The positive clone exhibiting a white color phenotype was isolated and recombinant plasmid was sequenced.
 | |
 | |
| Fig. 5: | Alignment of amino acid sequences of amylase from various B. stearothermophilus species (ac: Y17557, X59476, AF032864, X02769, AY705090, M11450 and M57457) compared to YN amylase sequence. The alignment was performed with CLUSTAL W as described under material and methods |
The sequencing results presented in Fig. 4 revealed two ORFs, the first was a unique (GTG) with a molecular size 1649 nucleotides encoding 549aa residues of a predicted molecular weight 62.592 kD and the second (ATG) with a molecular size 1613 nucleotides encoding 537aa residues of a predicted molecular weight 61.04 kD, that showed 99% similarity to α-amylase from other thermophilic bacilli and geobacilli. Multialignment between the amino acid sequences of Geobacillus thermoleovorans YN and the closely related sequences (Geobacillus stearothermophilus, ac: Y17557, X59476, AF032864, X02769, AY705090, M11450 and M57457) showed minor differences in aa (Fig. 5), this reflects the very close relations between amylases from thermophiles. Also, this comparison has revealed that YN amylase showed high aa identity with α-amylases from Bacillus stearothermophilus. As explained by Ben Ali et al. (2001), despite there are minor aa changes between different B. stearothermophilus amylases, it can establish some important differences on either the end product of starch hydrolysis or on the thermostability and thermoactivity. Therefore there are a correlation between these aa changes and the physico-chemical properties of the enzyme.
Accordingly the study will attend to express the gene successfully and intensively characterize the produced YN amylase enzyme to find new applications. The scaling up using a well characterized enzyme produced by a recombinant construct will be more applicable and economic.
Geobacillus thermoleovorans YN constitutively produces a thermoactive α-amylase specifically the maltohexaose-α-amylase as explained with sequence similarity analysis. The present study addressed the production of this enzyme by this bacterial cell in different media and zymography of the crude extract which revealed the presence of one active band of apparent molecular weight around 62 kD was performed. This matched with the predicted molecular weight of the ORF (GTG) of the revealed gene sequences. It is worthwhile to express this gene under a defined promoter using all the probable ORFs individually.