Research Article
Isolation and Characterization of a Yellow Pigmented Colony Forming Bacterium for Carotenogenesis
Tamil Nadu Agriculture University, India
LiveDNA: 91.33845
V. Udayasuriyan
Tamil Nadu Agriculture University, India
Carotenoids are yellow, orange and red pigments, which are widely distributed in nature (Goodwin and Britton, 1988). In microorganisms and plants where carotenoids are synthesized, their major function is protection against oxidative damage. These pigments are capable of quenching photosensitizers, interacting with singlet oxygen (Krinsky, 1994) and scavenging peroxy radicals (Conn et al., 1992). Industrially, carotenoid pigments are utilized as food colorants and feed supplements. Recently, carotenoids have attracted greater attention due to their beneficial effect on human health such as involvement in cancer prevention (Smith, 1998; Guerin et al., 2003) and enhancement of immune response (Johnson, 2002). β-carotene is the precursor for vitamin A. Deficiency of vitamin A is recognized as a serious public health problem, which contributes to a considerable proportion of blindness in children (Rajalakshmi et al., 2001).
In recent years, cloning of carotenogenic (crt) genes from various sources has made it possible to synthesize carotenoids in non-carotenogenic microorganisms and plants using recombinant DNA technology (Yokoyama et al., 1998; Mann et al., 2000). The phytoene desaturase (crtI) gene from Pantoea annanatis was used in the engineering of carotenoid biosynthetic pathway in rice (Ye et al., 2000). The present study describes identification of crtI and crtB genes in a new isolate of Pantoea sp. Identification of new crt genes in indigenous isolates of bacteria would provide the source to produce carotenoids in heterologous hosts.
The present study was conducted at the Department of Biotechnology, Centre for Plant Molecular Biology, Tamil Nadu Agricultural University, Coimbatore during 2004. The yellow-pigmented colony forming bacterial strain C1B1Y was isolated from the surroundings of our laboratory, Coimbatore, India, by air sampling. The Luria-Bertani growth medium containing (1 g-1): Tryptone 10; Sodium chloride 10; Yeast extract 5 and pH 7.0 was used. The strain was pure cultured by single colony isolation and microscopic analysis (1000x) was done after Gram staining (Baron et al., 1995). The bacterial isolate C1B1Y was grown aerobically in LB broth in a rotary shaker at 180 rpm at 30°C. After three days, cells were harvested by centrifugation (4000 rpm, 10 min, 4°C), washed with distilled water and the pigment was extracted from the pellet with acetone at 60°C for 20 min or until all visible pigments were extracted. Wet cells from 40 mL culture were extracted with 6 mL of methanol. After centrifugation (4000 rpm, 10 min, 4°C) the coloured supernatant was separated and filtered. For the analysis of carotenoids the acetone extract was monitored from 300 to 600 nm in a Varian Cary 50 Conc UV-Visible spectrophotometer. Analysis of C1B1Y pigment extract by HPLC was done using Shimadzu LC-8A with photo diode array detector. Acetonitrile was used as mobile phase with a flow rate of 1 mL min-1 and 18C column (Phenomenex, 250x4.6 mm, 5 μm) was used. The β-carotene from Himedia, India was used as reference sample for absorption spectrum and HPLC analysis.
Total genomic DNA was isolated from the bacterial isolate C1B1Y using GenElute bacterial genomic DNA kit, Sigma. Amplification of 16S rDNA region by PCR was performed with 27f and 519r primers (Lane, 1991). The 16S rDNA amplicon was cloned into the vector pTZ57R, (InsT/A clone PCR product cloning kit, MBI Fermentas, Lithuania) and sequenced through M13 forward and reverse primers. Automated DNA sequencing service (Bangalore Genei, India) was used for sequencing. The partial 16S rDNA sequence of C1B1Y was analysed by Ribosome Database Project (RDP-II)-Sequence Match for comparison. The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain C1B1Y is AY876937.
The total genomic DNA of bacterial isolate C1B1Y was screened by PCR with degenerate primers specific to the conserved region of crtI and crtB genes (Hannibal et al., 2000). A touchdown PCR (Don et al., 1991) was done as follows: initial denaturation at 94°C for 5 min followed by 20 cycles consisting of a 30 sec denaturation at 94°C, 30 sec at an annealing temperature of 60 to 50°C and a 1 min primer extension at 72°C, followed by 15 cycles consisting of a 30 sec denaturation at 94°C, 30 sec at an annealing temperature at 50°C and a 1 min primer extension at 72°C. The amplified DNA fragment of C1B1Y was analysed in 0.8% agarose gel and the eluted DNA fragment of C1B1Y was ligated into the vector pTZ57R (2.88 kb). The ligated DNA was used to transform DH5α E. coli strain. Recombinant clones were identified by blue/white colony selection.
Plasmid isolated from two recombinant clones were used separately for automated DNA sequencing. The crtI-crtB sequence data of C1B1Y was compared with representative sequences of Pantoea sp. obtained from the GenBank. Deduced amino acid sequence, was obtained using the Bioedit sequence alignment editor program version 5.0.9 (Hall, 1999). Multiple sequence alignment of the amino acid sequences was performed using ClustalX (Thompson et al., 1997). The nucleotide sequence of the 612 bp crtI-crtB region has been deposited in the GenBank/EMBL/DDBJ database under accession No. AY876938.
A yellow-pigmented bacterial colony (C1B1Y) was isolated by air sampling. Microscopic observation of the yellow colony forming strain C1B1Y proved it to be Gram-negative, rod shaped bacterium.
 | |
| Fig. 1: | Absorption spectra of the acetone extract (a) β-carotene standard (λmax = 449.9 nm) (b) Bacterial isolate C1B1Y (λmax = 450 nm) |
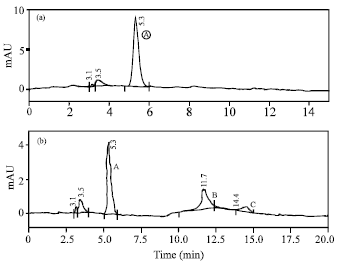 | |
| Fig. 2: | HPLC analysis of pigments extracted from the β-carotene standard; (a) the bacterial isolate C1B1Y ; (b) chromatograms were recorded as a function of the absorbance (mAU, milliabsorbance units) between 300 and 600 nm. Peak identities: Peak A-β-carotene, peak B and C-unidentified |
All pigmented bacterial isolates need not be carotenogenic (Moss, 2002). With very few exceptions, carotenoids are lipophilic. They are insoluble in water and are soluble in organic solvents. The spectrum analysis of organic solvent extract is the first diagnostic tool for the identification of carotenoids (Rodriguez-Amaya, 1999).
 | |
| Fig. 3: | Amplification of the crtI-crtB region of the bacterial isolate C1B1Y by PCR based on the degenerate primers specific for the conserved regions of crtI and crtB. Lane 1: 100 bp marker, Lane 2: Amplicon from the C1B1Y isolate |
The absorption spectrum of acetone extract of the cell pellet from the bacterial isolate C1B1Y showed absorption maximum of 450 nm which was identical to the absorption spectrum of the β-carotene reference sample (Fig. 1). The Absorption maximum of acetone extract of C1B1Y strain is also very close to that of the standard β-carotene in acetone (452 nm) (Rodriguez-Amaya, 1999). Moreover, the retention time in HPLC analysis of both the β-carotene standard and the extract of C1B1Y were identical (Fig. 2). These results conclude the production of β-carotene by the new isolate C1B1Y.
Comparison of the partial 16S rDNA sequence (486 bp) of C1B1Y strain by the Ribosomal Database Project II-Sequence match revealed the greatest sequence identity with Pantoea agglomerans (GenBank accession no U80202; S_ab = 0.957). This leads us to suggest that the C1B1Y isolate is more related to P. agglomerans.
 | |
| Fig. 4: | Alignment of deduced amino acid sequence of C1B1Y crtI-crtB region with corresponding region of Pantoea sp. generated by ClustalX. Residues 1-29 represent C terminal end of crtI and residues 31-205 represent N terminal end of crtB. The crtI-crtB region of C1B1Y shows variation in three amino acid residues, at positions 26, 58 and 198 on comparison with its nearest homolog of P. agglomerans (AB076662). Peptide numbering refers to the position in the cloned PCR fragment rather than to that of the full-length enzyme. The GenBank accession number details are as follows: Pantoea agglomerans (AB076662, M90698, M87280 and M38423), Pantoea ananatis (D90087) and Pantoea stewartii (AY166713) |
Carotenogenic genes (crt) in bacteria are known to be present in a cluster. The genes for phytoene desaturase (crtI) and phytoene synthase (crtB) are known to be present in sequence. The indigenous bacterial isolate C1B1Y was screened for the presence of crtI and crtB genes using degenerate primers corresponding to conserved regions of crtI and crtB. The bacterial isolate C1B1Y gave amplification of about 620 bp (Fig. 3). Nucleotide sequence analysis of the DNA fragment (612 bp) cloned from the crtI-crtB region of the new isolate C1B1Y showed highest homology (98%) with that of P. agglomerans (GenBank accession no AB076662). Comparison of deduced amino acid sequence of the C1B1Y isolate with its nearest homolog P. agglomerans (GenBank accession no AB076662) revealed variation of three amino acid residues between the conserved region of the CrtI and CrtB (Fig. 4). This indicates presence of crtI and crtB genes in the new isolate of Pantoea sp. (C1B1Y).
We thank P. Sangeetha, Senior Research Fellow, for her assistance during the isolation of yellow-pigmented bacteria. We thank Dr. R. Samiyappan, Professor, Department of Plant Pathology, Tamil Nadu Agricultural University, Coimbatore and Dr. K. P. Salin, Senior Scientist, Section of Entomology, Sugarcane Breeding Institute, Coimbatore for kindly providing the spectrophotometer and HPLC facilities, respectively.