ABSTRACT
In this study, karyotype analyses of the three species of Cicer L. were examined. Chromosome lengths were determined between 2.18-2.87 μm in Cicer bijugum Rech. f., 2.74-4.24 μm in Cicer oxyodon Boiss. and Hohen, 2.14-3.02 μm in Cicer anatolicum Alef., respectively. All species were diploid with 2n = 2x = 16 chromosomes and karyotypes of all species consist of metacentric chromosome pairs. Cicer oxyodon had only one satellite on the short arm of chromosome 7. The karyotype analysis of these species was first time counted in Iran.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajcb.2011.97.104
URL: https://scialert.net/abstract/?doi=ajcb.2011.97.104
INTRODUCTION
Genus Cicer L. (Fabaceae) of tribe Cicereae Alef., is best known by the globally important cultivated species C. arietinum L., other than the cultivated species, it is known to have about 10 annuals and 34 perennial relatives. About 40 species grow in Middle and West Asia and about 4 species in certain regions of north and northwestern areas of Africa and Europe (Van der Maesen, 1987; Ahmad, 2000).
This genus consists of some herbaceous and shrubby species which are classified into 4 sections (Monocicer, chamaecicer, polycicer and Achanthocicer) on the basis of morphological specifications and life cycle traits (Van der Maesen, 1987, 1972). The understudied Polycicer species were belongs to Monocicer (C. bijugum, annual species) and Polycicer (C. oxyodon and C. anatolicum, perennial species) sections.
The section, Monocicer comprising all annual species is most important to breeders. The related wild species are an immense reservoir of useful genetic traits and attempts are already in progress to exploit it for the genetic improvement of cultivated chickpea (Singh et al., 1998). C. bijugum used in the crossing program has many desirable characters such as resistance to Ascochyta blight, botrytis grey mold and to Helicoverpa-the menacing pod borer.
Chromosome number in Cicer species can be generalized as 2n = 2x = 16, although varying numbers both for chickpea (2n = 2x = 14, 16, 24, 32, 33) and other wild Cicer species (2n = 14, 16, 24) have been reported but could not be confirmed by other workers (Iyengar, 1939; Singh, 1964; Sohoo et al., 1970; Mercys et al., 1974; Polhill and Van Der Maesen, 1985; Tayyar et al., 1994). The 2n chromosomes number of Cicer bijugum and Cicer anatolicum has been reported to be 16 (Ahmad, 1989; Ahmad and Chen, 2000). There was no report to date on the detailed karyotype of these species. Thus the karyotype analysis of these species was first time counted in Iran. This information should be useful in determining phylogenetic relationships among the different species of the genus Cicer and also this study was undertaken with a view to confirm the chromosome number of the different species.
MATERIAL AND METHODS
The materials used in this study were collected in different areas of Iran. The localities, gene bank codes and species names are shown in Table 1. Vouchers are deposited in gene bank RIFR (Research Institute of Forest and Rangelands from Iran). Root tip meristems obtained from seedlings were pretreated with 0.5% saturated α-Bromo naphthalene at 4°C for 4 h, fixed in 10% formaldehyde and 1% chromic acid (1:1) for at least 16 h at room temperature, then root tips were rinsed for 3 h in distilled water. Hydrolysis was carried out with 1 N NaOH at 60°C for 6 min, dyed with hematoxylin for 3-4 h and squashed in a droplet of 45% acetic acid and lactic acid (10:1). The preparations were observed with an optical microscope (BH2 Olympus supplemented digital color video camera) at a magnification of 1908x.
The best metaphysical plates were selected and measured by Micro measure 3.3 software (Reeves and Tear, 2000). In each mitotic metaphase (at least 5 plates) the arm's length of each chromosome was measured, according to the previous studies (Hesamzaden Hejazi and Rasuli, 2006; Hesamzadeh and Ziaei Nasab, 2009, 2010).
The following parameters were estimated in each metaphase plate to characterize the karyotypes numerically: Long Arm (LA), Short Arm (SA), Total Length (TL) [LA+SA], Relative Length percentage (RL%), Arm Ratio (AR) [LA/SA], Centromeric Index (CI) [SA/ (LA+SA)], Value of Relative Chromatin (VRC) [ΣTL/n]. Karyotype asymmetry was estimated by three different methods namely, Total Form percentage (TF%) [(ΣSA/ΣTL) x100] (Huziwara, 1962); Difference of Relative Length (DRL) [MaxRL% - MinRL%]; Dispersion Index (DI)[%CVxCG] where CV represents the coefficient of variation for chromosome length and CG represents the centromeric gradient value [(length of median short arm /length of median chromosome)x100] (Lavania and Srivastava, 1992); %SI; intrachromosomal asymmetry index (A1) [1-Σ (![]() )/n] and interchromosomal asymmetry index (A2) [
)/n] and interchromosomal asymmetry index (A2) [![]() ] (Zarco, 1986). Both indices (A1 and A2) are independent to chromosome number and size.
] (Zarco, 1986). Both indices (A1 and A2) are independent to chromosome number and size.
Also karyotypic evolution has been determined using the symmetry classes of Stebbins (SC) Stebbins (1971). Karyotype formula was determined by chromosome morphology based on centromere position according to classification of Levan et al. (1964). For each species, karyograms and haploid idiograms were drawn based on length of chromosome size (arranged large to small).
In order to determine the variation between species, one-way balanced ANOVA was performed on normal data and parameter means were compared by Duncan’s test. Numerical analyses were performed using SAS (1996) software ver. 6.12.
RESULTS
Karyotype analyses of the three species of Cicer were determined. All somatic cells of investigated species have diploid chromosomes with 2n = 16. Average of chromosomal length of investigated species ranged from 2.46 to 3.43 μm.
| Table 1: | Localities of species used in the study |
 | |
| Table 2: | Measurements (μm) of somatic metaphase chromosomes of C. oxyodon |
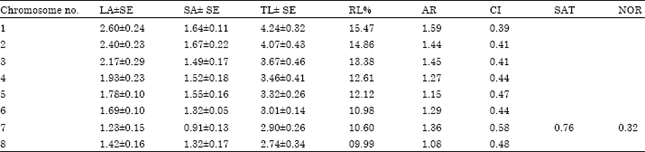 | |
| Long arm (LA), short arm (SA), total length (TL), relative length percentage (RL%), arm ratio (AR), centromeric index (CI),Satellite(SAT), Nucleolar Organizer Region (NOR) | |
| Table 3: | Measurements (μm) of somatic metaphase chromosomes of C. bijugum |
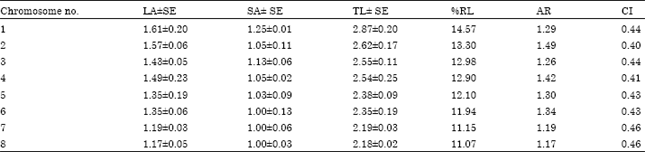 | |
| Long arm (LA), short arm (SA), total length (TL), relative length percentage (RL%), arm ratio (AR), centromeric index (CI),Satellite(SAT), Nucleolar Organizer Region (NOR) | |
| Table 4: | Measurements (μm) of somatic metaphase chromosomes of C. anatolicum |
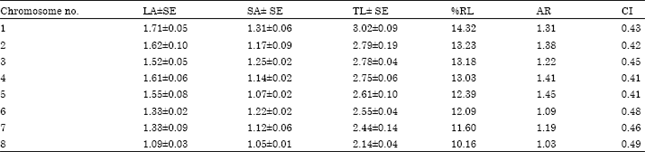 | |
| Long arm (LA), short arm (SA), total length (TL), relative length percentage (RL%), arm ratio (AR), centromeric index (CI),Satellite(SAT), Nucleolar Organizer Region (NOR) | |
Among these, while C. anatolicum has the smallest chromosome (Fig. 3), the biggest chromosome was observed on C. oxyodon (Fig. 1). Detailed measurements of somatic chromosomes of three Cicer species are given in Table 2-5. Among these, C. oxyodon has bigger haploid complement than other species.
Mean length of the haploid chromosome complement, recorded from at least five cells, were 27.41±0.19 μm, 19.68±0.08 μm, 21.08±0.09 μm respectively in C. oxyodon, C. bijugum and C. anatolicum.
Based on total length, chromosome pairs were arranged from 1 to 8 in order of decreasing length (Fig. 1-3). Chromosome length of the complement ranged from 2.74 to 4.24 μm, with a mean chromosome size of 3.43±0.18 μm observed in C. oxyodon. Based on the location of the primary constriction and secondary constriction, the following karyotypic formula was developed for C. oxyodon: K.F.(n = 16) = 16 m with a pair small satellite connected to the short arm of the seventh chromosome.
| Table 5: | Karyotype characteristics of three species of Cicer |
 | |
| 2n: somatic chromosome number, SC: Symmetry classes of Stebbins, A1: Intrachromosome asymmetry index, A2: Interchromosome asymmetry index, TF%: Total Form percentage, DRL: Difference of Relative Length, VRC: Value of Relative Chromatin, SI: Symmetry Index percentage, DI: Dispersion Index, K.F.: Karyotype Formulae (m: metacentric) | |
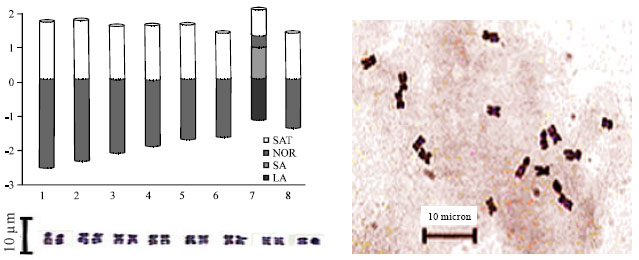 | |
| Fig. 1: | Metaphase plates, idiogram and karyogram of somatic chromosomes of C. oxyodon |
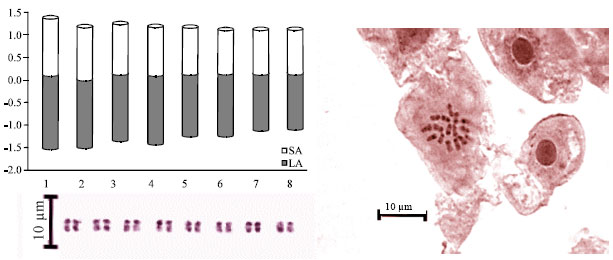 | |
| Fig. 2: | Metaphase plates, Idiogram and karyogram of somatic chromosomes of C. bijugum |
Chromosome length of the complement ranged from 2.18 to 2.87 μm, with a mean chromosome size of 2.46±0.08 μm observed in C. bijugum. Based on the location of the primary constriction the following karyotypic formula was developed for C. bijugum: K.F. (n = 16) = 16 m. Chromosome length of the complement ranged from 2.14 to 3.02 μm, with a mean chromosome size of 2.64±0.09 μm observed in C. anatolicum. Based on the location of the primary constriction the following karyotypic formula was developed for C. anatolicum: K.F. (n = 16) = 16 m.
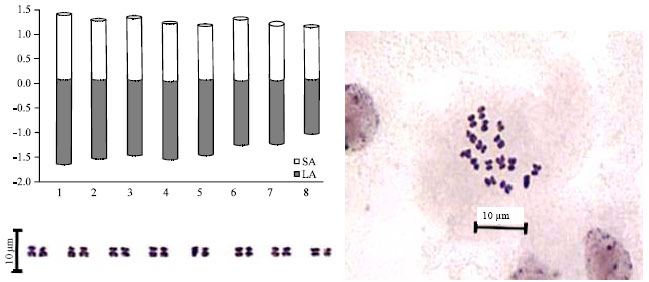 | |
| Fig. 3: | Metaphase plates, Idiogram and karyogram of somatic chromosomes of C. anatolicum |
| Table 6: | The results of analysis of variance for karyotypic data based on CRD design |
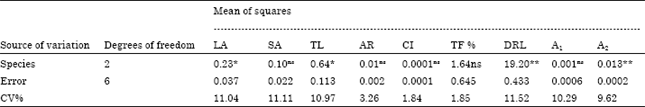 | |
| *- Significant at 5% level of probability, **- Significant at 1% level of probability, ns- Non significant | |
| Table 7: | Mean of chromosomes analysis of Cicer species |
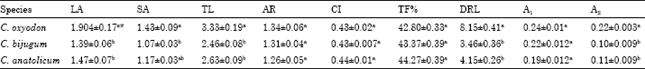 | |
| LA:long arm, SA: short arm, TL: total length, AR: arm ratio, CI: centromic index, TF%: Total Form percentage, DRL: Difference of Relative Length, A1: intrachromosome asymmetry index, A2: interchromosome asymmetry index, * indicated Mean within each column followed by different letters are significantly different at the 5% level according to the Duncan's test | |
Fundamentally, all of the studied species showed very similar karyotypic characters except on TL, LA, DRL and A2 traits. The dispersion index is calculated as the proportionate measure of centromeric gradient to the coefficient of variation for chromosome length. The highest value of DI was found in C. anatolicum (11.54) and the lowest value of DI was found in C. bijugum (3.81) species.
Symmetry type of Stebbins (1971) and asymmetry indices of Zarco (1986) are given in (Table 5).
A statistical comparison based on completely randomized design demonstrates that there are nonsignificant differences among the species for all the measured traits except, TL and LA traits with (p<5%) and DRL and A2 traits with (p<1%) (Table 6).
The Duncan’s test applied to the chromosome morphometric traits showed a non significant difference among the three species, except for TL, LA, DRL and A2 traits (Table 7).
DISCUSSION
In this study, chromosome numbers and detailed measurements of three species of Cicer genus were determined for the first time in Iran. A number of investigators have examined the chromosome numbers of the Cicer genus. However, out of the 24 different species listed in Flora Orientalis (Boissier, 1872) and Index Kewensis (Hooker and Jackson, 1895), the chromosome numbers of only 4 species have been studied. All the so far studied Cicer species have revealed a diploid chromosome numbers of 2n = 14 and 16.
The karyotype of the cultivated chickpea, Cicer arietinum, has already been reported (Sharma and Gupta, 1982; Ahmad and Scoles, 1987).
Generally, the karyotype concept has been extensively used in characterizing and distinguishing chromosomes of different species. Mitotic karyotype analyses are also helpful in studying evolutionary problems (Gottschalk, 1972).
The results showed only minor differences in gross morphology of the karyotypes of C. oxyodon, C. bijugum and C. anatolicum species. In order to refine the measure of karyotype asymmetry, we used Dispersion Index (DI) that has the potential to decipher even the minor karyotypic variations. The DI index plays an important role in arranging the species within the same class of karyotype asymmetry in an advancing order of specialization by permitting further gradations, as depicted by species arrangement within sections. Higher values of DI index would mean an enhanced order of karyotypic specialization.
Karyotype of C. bijugum is characterized by symmetrical karyotypes and comparatively smaller mean haploid chromosome complement lengths and mean chromosome sizes.
A well defined pair of satellites chromosomes was clearly distinguishable in C. oxyodon species. The secondary constriction (nucleolar organizing region) in C. oxyodon species was present on chromosome 7 about 0.32 μm from the primary constriction and was quite conspicuous in both the homologous of all the cells observed. In C. oxyodon, mean total length, lengths of long arm and short arm chromosomes were more than other species.
Only significant effect of TL, LA and DRL or A2 among species proved that the three species have karyotypes that resemble very closely to each other except on total length of chromosomes.
In view of the fact that, fewer DRL values illustrated more symmetry of karyotype, C. bijugum and C. oxyodon with DRL = 3.46 and 8.15 have the most symmetric and asymmetric karyotypes respectively. Similarly, High DRL value leads to more changes in the construction of chromosomes, therefore there must be more changes in C. oxyodon chromosomal construction.
In this study karyotype analyses of C. anatolicum, have given eight metacentric pairs of chromosomes but this result disagree with other research on same species (Ahmad, 1989).
In conclusion, from the examination of the chromosome numbers of the different species of Cicer it seems logical to conclude that the diploid number of chromosomes for the genus Cicer is 16. With regard to the number of pairs of satellites chromosomes (Iyengar, 1939) has reported that some species of Cicer have two pairs of satellites chromosomes. In the present investigation, only one pair of satellites chromosome was observed just in C. oxyodon.
Such karyotypic similarities and cross ability/genetic relationships raise an important curiosity as to whether there is any valid relationship between karyotypic similarity and cross ability in the genus Cicer and if this could give us any insight into genetic relationships. According to Lamprecht (1984) closely related species differ genetically from one another with regard to one or a few genes and thus a genetic barrier between the species can exist without cytological differences. Thus, it is important that other Cicer species also be studied to gain further understanding regarding genetic relationships and phylogeny in the genus Cicer. Crossing the cultivated and wild chickpea is expected to produce a hardy plant that will be able to stand up better to harsh weather and pest attacks.
REFERENCES
- Ahmad, F. and G.J. Scoles, 1987. The cytogenetic relationship between Cicer judaicum Boiss and Cicer chorassanicum (Bge) M. Pop Genome, 29: 883-886.
Direct Link - Huziwara, Y., 1962. Karyotype analysis in some genera of compositae. V III, Further studies on the chromosome of aster. Am. J. Bot., 49: 116-119.
Direct Link - Iyengar, N.K., 1939. Cytological investigations on the genus Cicer. Ann. Bot., 3: 271-305.
Direct Link - Mercy, T., S.K. Kakar and J.B. Chowdhry, 1974. Cytological studies in three species of the genus Cicer. Cytologia, 39: 383-390.
Direct Link - Zarco, C.R., 1986. A new method for estimating karyotype asymmetry. Taxon, 35: 526-530.
CrossRefDirect Link - Sohoo, M.S., D.S. Athwalet and S. Chandra, 1970. Colchicine induced polyploidy in chickpeas (Cicer arietinum L.). Theoret. Applied Genet., 40: 163-168.
CrossRefDirect Link - Stebbins, G.L., 1971. Chromosomal Evolution in Higher Plants. 1st Edn., Edward Arnold Publisher Ltd., London, UK., Pages: 216.
Direct Link - Tayyar, R.I., A.J. Lukaszewski and J.G. Waines, 1994. Chromosome banding patterns in the annual species of Cicer. Genome, 37: 656-663.
CrossRefPubMedDirect Link - Hesamzadeh, H.S.M. and M. Rasouli, 2006. Cytogenetic study of some species of Vetch Genus (Vicia sp.) in Iran. Iran. J. Agric. Sci., 37: 213-225.
Direct Link - Hesamzadeh, H.S.M. and M. ZiaeiNasab, 2009. Cytogenetic study on several populations of diploid species of Onobrychis in natural gene bank of Iran. Iran. J. Rangelands Forests Plant Breed. Genet. Res., 16: 158-171.
Direct Link - Hesamzadeh, H.S.M. and M. Ziaei Nasab, 2010. Cytotaxonomy of some Onobrychis (Fabaceae) species and populations in Iran. Caryologia, 63: 18-31.
Direct Link - Lavania, U.C. and S. Srivastava, 1992. A simple parameter of dispersion index that serves as an adjunct to karyotype asymmetry. J. Biosci., 17: 179-182.
CrossRefDirect Link - Singh, K.B., B. Ocampo and L.D. Robertson, 1998. Diversity for abiotic and biotic stress resistance in the wild annual Cicer species. Genet. Resour. Crop Evol., 45: 9-17.
CrossRef









Efat Rajaeian Reply
I would like to thank the management of this Journal and also the author that thay prepare this paper.
this paper realy was good for our porpose.